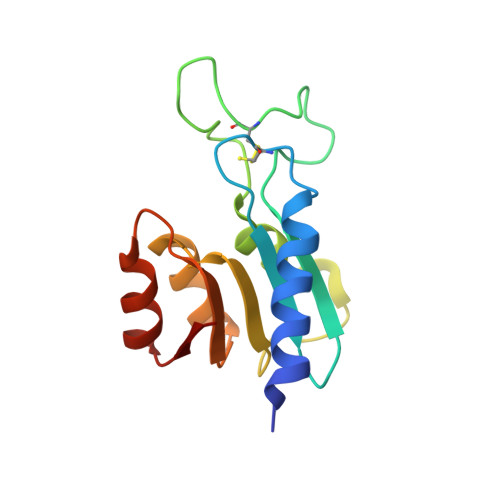
The First Crystal Structure of a dTTP-bound Deoxycytidylate Deaminase Validates and Details the Allosteric-Inhibitor Binding Site.
Marx, A., Alian, A.(2015) J Biological Chem 290: 682-690
- PubMed: 25404739 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.617720
- Primary Citation Related Structures:
4P9C, 4P9D, 4P9E - PubMed Abstract:
Deoxycytidylate deaminase is unique within the zinc-dependent cytidine deaminase family as being allosterically regulated, activated by dCTP, and inhibited by dTTP. Here we present the first crystal structure of a dTTP-bound deoxycytidylate deaminase from the bacteriophage S-TIM5, confirming that this inhibitor binds to the same site as the dCTP activator. The molecular details of this structure, complemented by structures apo- and dCMP-bound, provide insights into the allosteric mechanism. Although the positioning of the nucleoside moiety of dTTP is almost identical to that previously described for dCTP, protonation of N3 in deoxythymidine and not deoxycytidine would facilitate hydrogen bonding of dTTP but not dCTP and may result in a higher affinity of dTTP to the allosteric site conferring its inhibitory activity. Further the functional group on C4 (O in dTTP and NH2 in dCTP) makes interactions with nonconserved protein residues preceding the allosteric motif, and the relative strength of binding to these residues appears to correspond to the potency of dTTP inhibition. The active sites of these structures are also uniquely occupied by dTMP and dCMP resolving aspects of substrate specificity. The methyl group of dTMP apparently clashes with a highly conserved tyrosine residue, preventing the formation of a correct base stacking shown to be imperative for deamination activity. The relevance of these findings to the wider zinc-dependent cytidine deaminase family is also discussed.
- From the Faculty of Biology, Technion-Israel Institute of Technology, Haifa 320003, Israel.
Organizational Affiliation: