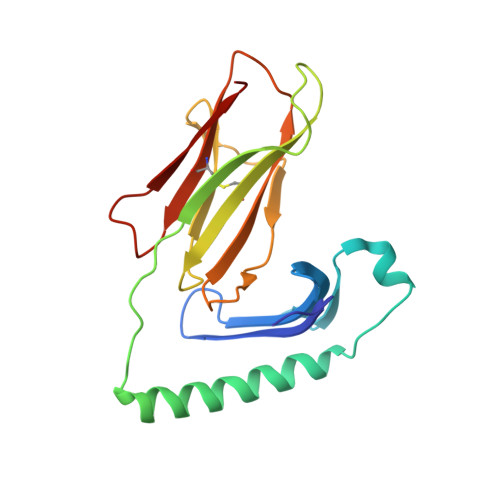
Structural basis of chronic beryllium disease: linking allergic hypersensitivity and autoimmunity.
Clayton, G.M., Wang, Y., Crawford, F., Novikov, A., Wimberly, B.T., Kieft, J.S., Falta, M.T., Bowerman, N.A., Marrack, P., Fontenot, A.P., Dai, S., Kappler, J.W.(2014) Cell 158: 132-142
- PubMed: 24995984
- DOI: https://doi.org/10.1016/j.cell.2014.04.048
- Primary Citation of Related Structures:
4P4K, 4P4R, 4P57, 4P5K, 4P5M - PubMed Abstract:
T-cell-mediated hypersensitivity to metal cations is common in humans. How the T cell antigen receptor (TCR) recognizes these cations bound to a major histocompatibility complex (MHC) protein and self-peptide is unknown. Individuals carrying the MHCII allele, HLA-DP2, are at risk for chronic beryllium disease (CBD), a debilitating inflammatory lung condition caused by the reaction of CD4 T cells to inhaled beryllium. Here, we show that the T cell ligand is created when a Be(2+) cation becomes buried in an HLA-DP2/peptide complex, where it is coordinated by both MHC and peptide acidic amino acids. Surprisingly, the TCR does not interact with the Be(2+) itself, but rather with surface changes induced by the firmly bound Be(2+) and an accompanying Na(+) cation. Thus, CBD, by creating a new antigen by indirectly modifying the structure of preexisting self MHC-peptide complex, lies on the border between allergic hypersensitivity and autoimmunity.
- Howard Hughes Medical Institute, National Jewish Health, Denver, CO 80206, USA; Department of Biomedical Research, National Jewish Health, Denver, CO 80206, USA; Department of Immunology and Microbiology, University of Colorado Denver School of Medicine, Aurora, CO 80045, USA.
Organizational Affiliation: