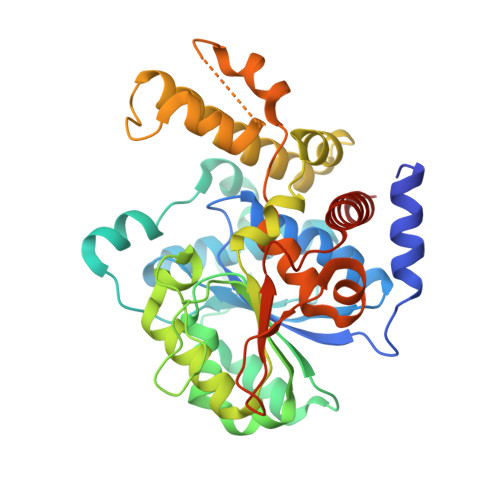
The crystal structure of type III effector protein XopQ from Xanthomonas oryzae complexed with adenosine diphosphate ribose.
Yu, S., Hwang, I., Rhee, S.(2014) Proteins 82: 2910-2914
- PubMed: 25079351 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24656
- Primary Citation Related Structures:
4P5F - PubMed Abstract:
Effector proteins are virulence factors that promote pathogenesis by interfering with various cellular events and are delivered directly into host cells by the secretion systems of many Gram-negative bacteria. Type III effector protein XOO4466 from the plant pathogen Xanthomonas oryzae pv. oryzae (XopQ(Xoo)) and XopQ homologs from other phytopathogens have been predicted to be nucleoside hydrolases based on their sequence similarities. However, despite such similarities, recent structural and functional studies have revealed that XopQ(Xoo) does not exhibit the expected activity of a nucleoside hydrolase. On the basis of the conservation of a Ca(2+) coordination shell of a ribose-binding site and the spacious active site in XopQ(Xoo), we hypothesized that a novel compound containing a ribosyl moiety could serve as a substrate for XopQ(Xoo). Here, we report the crystal structure of XopQ(Xoo) in complex with adenosine diphosphate ribose (ADPR), which is involved in regulating cytoplasmic Ca(2+) concentrations in eukaryotic cells. ADPR is bound to the active site of XopQ(Xoo) with its ribosyl end tethered to the Ca(2+) coordination shell. The binding of ADPR is further stabilized by interactions mediated by hydrophobic residues that undergo ligand-induced conformational changes. These data showed that XopQ(Xoo) is capable of binding a novel chemical bearing a ribosyl moiety, thereby providing the first step toward understanding the functional role of XopQ(Xoo).
- Department of Agricultural Biotechnology, Seoul National University, Seoul, Korea.
Organizational Affiliation: