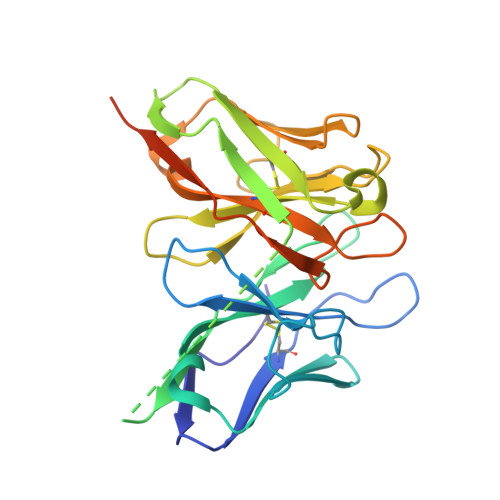
Reconciling the structural attributes of avian antibodies.
Conroy, P.J., Law, R.H., Gilgunn, S., Hearty, S., Caradoc-Davies, T.T., Lloyd, G., O'Kennedy, R.J., Whisstock, J.C.(2014) J Biological Chem 289: 15384-15392
- PubMed: 24737329 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.562470
- Primary Citation Related Structures:
4P48, 4P49 - PubMed Abstract:
Antibodies are high value therapeutic, diagnostic, biotechnological, and research tools. Combinatorial approaches to antibody discovery have facilitated access to unique antibodies by surpassing the diversity limitations of the natural repertoire, exploitation of immune repertoires from multiple species, and tailoring selections to isolate antibodies with desirable biophysical attributes. The V-gene repertoire of the chicken does not utilize highly diverse sequence and structures, which is in stark contrast to the mechanism employed by humans, mice, and primates. Recent exploitation of the avian immune system has generated high quality, high affinity antibodies to a wide range of antigens for a number of therapeutic, diagnostic and biotechnological applications. Furthermore, extensive examination of the amino acid characteristics of the chicken repertoire has provided significant insight into mechanisms employed by the avian immune system. A paucity of avian antibody crystal structures has limited our understanding of the structural consequences of these uniquely chicken features. This paper presents the crystal structure of two chicken single chain fragment variable (scFv) antibodies generated from large libraries by phage display against important human antigen targets, which capture two unique CDRL1 canonical classes in the presence and absence of a non-canonical disulfide constrained CDRH3. These structures cast light on the unique structural features of chicken antibodies and contribute further to our collective understanding of the unique mechanisms of diversity and biochemical attributes that render the chicken repertoire of particular value for antibody generation.
- From the Department of Biochemistry and Molecular Biology, Faculty of Medicine, Nursing and Health Science, Monash University, Melbourne, Victoria 3800, Australia.
Organizational Affiliation: