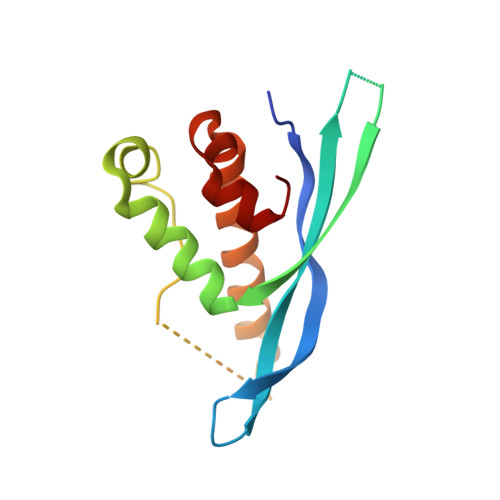
Structural Basis for Different Phosphoinositide Specificities of the PX Domains of Sorting Nexins Regulating G-protein Signaling.
Mas, C., Norwood, S.J., Bugarcic, A., Kinna, G., Leneva, N., Kovtun, O., Ghai, R., Ona Yanez, L.E., Davis, J.L., Teasdale, R.D., Collins, B.M.(2014) J Biological Chem 289: 28554-28568
- PubMed: 25148684 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.595959
- Primary Citation Related Structures:
4P2I, 4P2J, 4PQO, 4PQP - PubMed Abstract:
Sorting nexins (SNXs) or phox homology (PX) domain containing proteins are central regulators of cell trafficking and signaling. A subfamily of PX domain proteins possesses two unique PX-associated domains, as well as a regulator of G protein-coupled receptor signaling (RGS) domain that attenuates Gαs-coupled G protein-coupled receptor signaling. Here we delineate the structural organization of these RGS-PX proteins, revealing a protein family with a modular architecture that is conserved in all eukaryotes. The one exception to this is mammalian SNX19, which lacks the typical RGS structure but preserves all other domains. The PX domain is a sensor of membrane phosphoinositide lipids and we find that specific sequence alterations in the PX domains of the mammalian RGS-PX proteins, SNX13, SNX14, SNX19, and SNX25, confer differential phosphoinositide binding preferences. Although SNX13 and SNX19 PX domains bind the early endosomal lipid phosphatidylinositol 3-phosphate, SNX14 shows no membrane binding at all. Crystal structures of the SNX19 and SNX14 PX domains reveal key differences, with alterations in SNX14 leading to closure of the binding pocket to prevent phosphoinositide association. Our findings suggest a role for alternative membrane interactions in spatial control of RGS-PX proteins in cell signaling and trafficking.
- From the Institute for Molecular Bioscience, The University of Queensland, St. Lucia, Queensland 4072, Australia.
Organizational Affiliation: