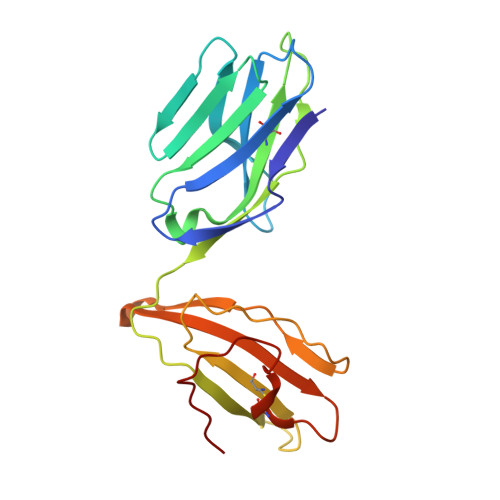
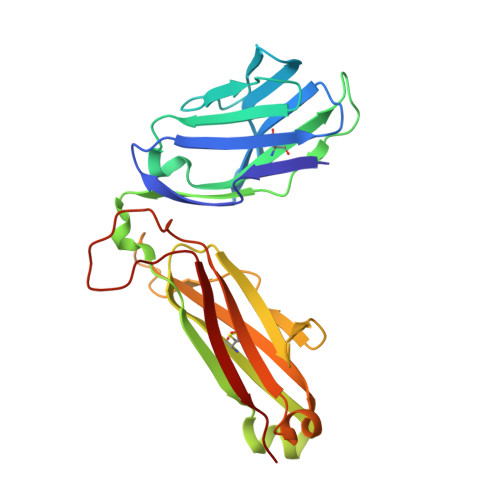
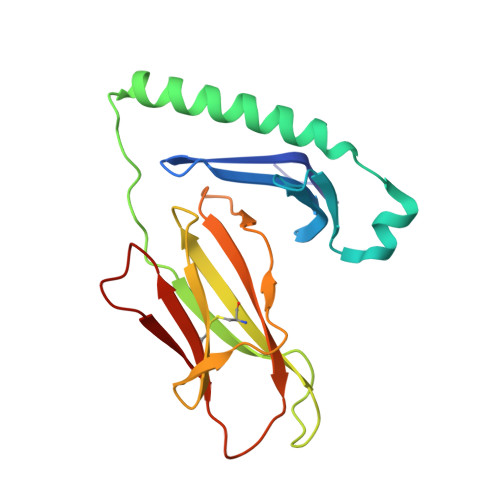
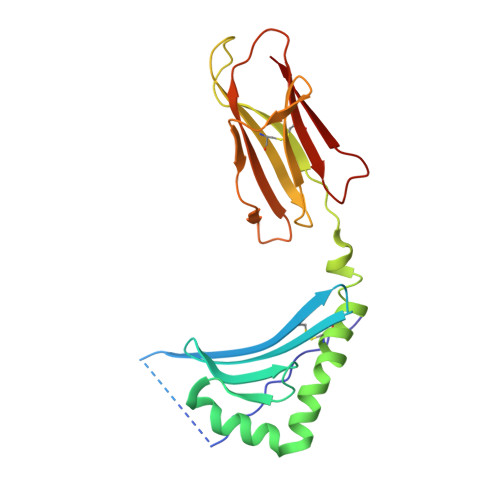
Effect of CDR3 Sequences and Distal V Gene Residues in Regulating TCR-MHC Contacts and Ligand Specificity.
Stadinski, B.D., Trenh, P., Duke, B., Huseby, P.G., Li, G., Stern, L.J., Huseby, E.S.(2014) J Immunol 192: 6071-6082
- PubMed: 24813203 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.4049/jimmunol.1303209
- Primary Citation Related Structures:
4P23, 4P46, 4P5T - PubMed Abstract:
The mature T cell repertoire has the ability to orchestrate immunity to a wide range of potential pathogen challenges. This ability stems from thymic development producing individual T cell clonotypes that express TCRs with unique patterns of Ag reactivity. The Ag specificity of TCRs is created from the combinatorial pairing of one of a set of germline encoded TCR Vα and Vβ gene segments with randomly created CDR3 sequences. How the amalgamation of germline encoded and randomly created TCR sequences results in Ag receptors with unique patterns of ligand specificity is not fully understood. Using cellular, biophysical, and structural analyses, we show that CDR3α residues can modulate the geometry in which TCRs bind peptide-MHC (pMHC), governing whether and how germline encoded TCR Vα and Vβ residues interact with MHC. In addition, a CDR1α residue that is positioned distal to the TCR-pMHC binding interface is shown to contribute to the peptide specificity of T cells. These findings demonstrate that the specificity of individual T cell clonotypes arises not only from TCR residues that create direct contacts with the pMHC, but also from a collection of indirect effects that modulate how TCR residues are used to bind pMHC.
- Department of Pathology, University of Massachusetts Medical School, Worcester, MA 01655; and.
Organizational Affiliation: