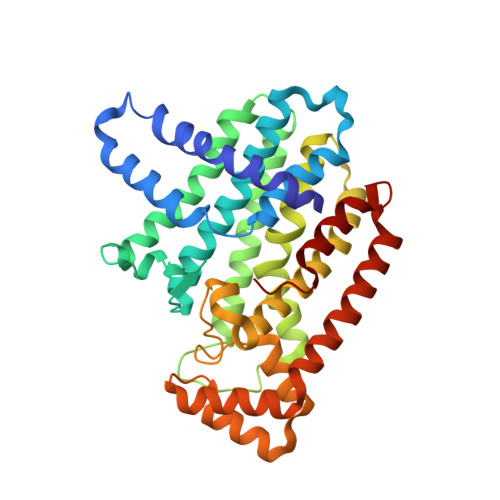
Taxodione and arenarone inhibit farnesyl diphosphate synthase by binding to the isopentenyl diphosphate site.
Liu, Y.L., Lindert, S., Zhu, W., Wang, K., McCammon, J.A., Oldfield, E.(2014) Proc Natl Acad Sci U S A 111: E2530-E2539
- PubMed: 24927548 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1409061111
- Primary Citation Related Structures:
4P0V, 4P0W, 4P0X - PubMed Abstract:
We used in silico methods to screen a library of 1,013 compounds for possible binding to the allosteric site in farnesyl diphosphate synthase (FPPS). Two of the 50 predicted hits had activity against either human FPPS (HsFPPS) or Trypanosoma brucei FPPS (TbFPPS), the most active being the quinone methide celastrol (IC50 versus TbFPPS ∼ 20 µM). Two rounds of similarity searching and activity testing then resulted in three leads that were active against HsFPPS with IC50 values in the range of ∼ 1-3 µM (as compared with ∼ 0.5 µM for the bisphosphonate inhibitor, zoledronate). The three leads were the quinone methides taxodone and taxodione and the quinone arenarone, compounds with known antibacterial and/or antitumor activity. We then obtained X-ray crystal structures of HsFPPS with taxodione+zoledronate, arenarone+zoledronate, and taxodione alone. In the zoledronate-containing structures, taxodione and arenarone bound solely to the homoallylic (isopentenyl diphosphate, IPP) site, not to the allosteric site, whereas zoledronate bound via Mg(2+) to the same site as seen in other bisphosphonate-containing structures. In the taxodione-alone structure, one taxodione bound to the same site as seen in the taxodione+zoledronate structure, but the second located to a more surface-exposed site. In differential scanning calorimetry experiments, taxodione and arenarone broadened the native-to-unfolded thermal transition (Tm), quite different to the large increases in ΔTm seen with biphosphonate inhibitors. The results identify new classes of FPPS inhibitors, diterpenoids and sesquiterpenoids, that bind to the IPP site and may be of interest as anticancer and antiinfective drug leads.
- Department of Chemistry and.
Organizational Affiliation: