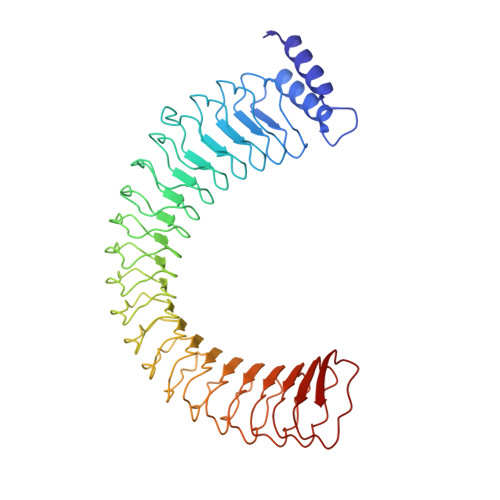
Immunosuppressive Yersinia Effector YopM Binds DEAD Box Helicase DDX3 to Control Ribosomal S6 Kinase in the Nucleus of Host Cells.
Berneking, L., Schnapp, M., Rumm, A., Trasak, C., Ruckdeschel, K., Alawi, M., Grundhoff, A., Kikhney, A.G., Koch-Nolte, F., Buck, F., Perbandt, M., Betzel, C., Svergun, D.I., Hentschke, M., Aepfelbacher, M.(2016) PLoS Pathog 12: e1005660-e1005660
- PubMed: 27300509 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1005660
- Primary Citation Related Structures:
4OW2 - PubMed Abstract:
Yersinia outer protein M (YopM) is a crucial immunosuppressive effector of the plaque agent Yersinia pestis and other pathogenic Yersinia species. YopM enters the nucleus of host cells but neither the mechanisms governing its nucleocytoplasmic shuttling nor its intranuclear activities are known. Here we identify the DEAD-box helicase 3 (DDX3) as a novel interaction partner of Y. enterocolitica YopM and present the three-dimensional structure of a YopM:DDX3 complex. Knockdown of DDX3 or inhibition of the exportin chromosomal maintenance 1 (CRM1) increased the nuclear level of YopM suggesting that YopM exploits DDX3 to exit the nucleus via the CRM1 export pathway. Increased nuclear YopM levels caused enhanced phosphorylation of Ribosomal S6 Kinase 1 (RSK1) in the nucleus. In Y. enterocolitica infected primary human macrophages YopM increased the level of Interleukin-10 (IL-10) mRNA and this effect required interaction of YopM with RSK and was enhanced by blocking YopM's nuclear export. We propose that the DDX3/CRM1 mediated nucleocytoplasmic shuttling of YopM determines the extent of phosphorylation of RSK in the nucleus to control transcription of immunosuppressive cytokines.
- Institute of Medical Microbiology, Virology and Hygiene, University Medical Center Hamburg-Eppendorf (UKE), Hamburg, Germany.
Organizational Affiliation: