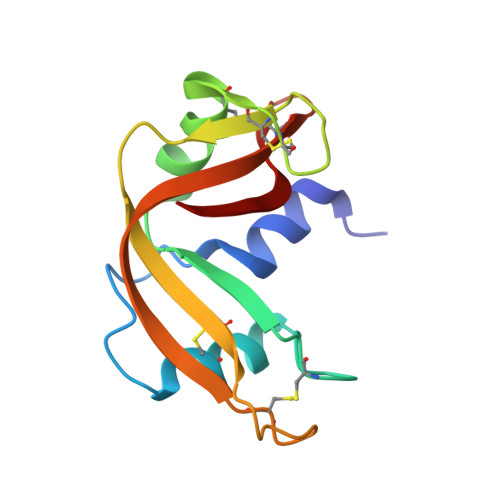
Cisplatin binding to proteins: molecular structure of the ribonuclease a adduct.
Messori, L., Merlino, A.(2014) Inorg Chem 53: 3929-3931
- PubMed: 24694179 Search on PubMed
- DOI: https://doi.org/10.1021/ic500360f
- Primary Citation Related Structures:
4OT4 - PubMed Abstract:
The crystal structure of the main adduct formed in the reaction between cisplatin and bovine pancreatic ribonuclease is reported here. Notably, in both of the protein molecules present in the asymmetric unit, platinum(II) binding takes place exclusively at the level of Met29. In one of the two molecules, the Gln28 side chain completes the platinum coordination sphere, anchoring the cisplatin fragment to the protein in a bidentate fashion. These results contain interesting implications for understanding the biological chemistry of this important drug.
- Department of Chemistry, University of Florence , Via della Lastruccia 3, 50019 Sesto Fiorentino, Italy.
Organizational Affiliation: