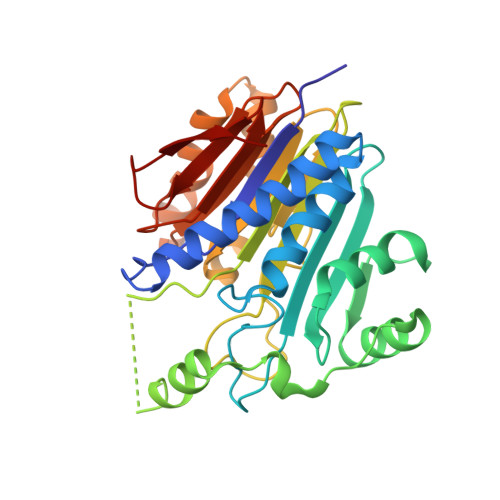
Free Glycine Accelerates the Autoproteolytic Activation of Human Asparaginase.
Su, Y., Karamitros, C.S., Nomme, J., Mcsorley, T., Konrad, M., Lavie, A.(2013) Chem Biol 20: 533
- PubMed: 23601642 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2013.03.006
- Primary Citation Related Structures:
4OSX, 4OSY - PubMed Abstract:
Human asparaginase 3 (hASNase3), which belongs to the N-terminal nucleophile hydrolase superfamily, is synthesized as a single polypeptide that is devoid of asparaginase activity. Intramolecular autoproteolytic processing releases the amino group of Thr168, a moiety required for catalyzing asparagine hydrolysis. Recombinant hASNase3 purifies as the uncleaved, asparaginase-inactive form and undergoes self-cleavage to the active form at a very slow rate. Here, we show that the free amino acid glycine selectively acts to accelerate hASNase3 cleavage both in vitro and in human cells. Other small amino acids such as alanine, serine, or the substrate asparagine are not capable of promoting autoproteolysis. Crystal structures of hASNase3 in complex with glycine in the uncleaved and cleaved enzyme states reveal the mechanism of glycine-accelerated posttranslational processing and explain why no other amino acid can substitute for glycine.
- Department of Biochemistry and Molecular Genetics, University of Illinois at Chicago, Chicago, IL 60607, USA.
Organizational Affiliation: