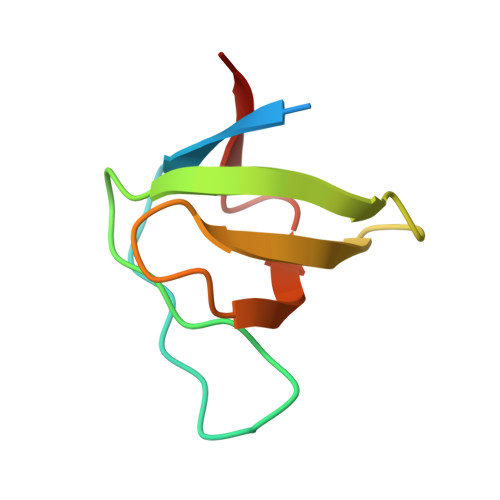
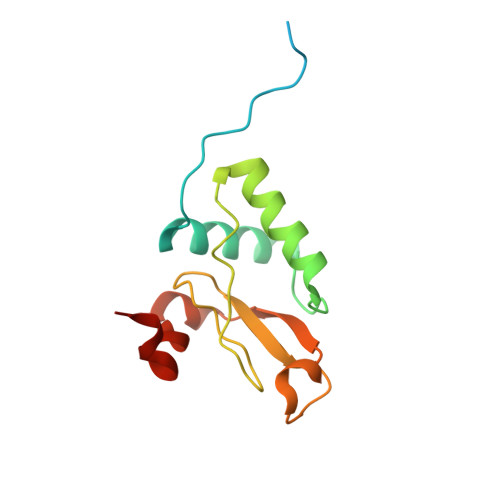
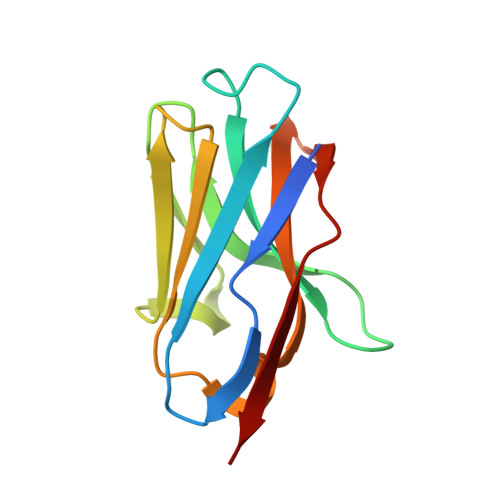
Structural basis for the inhibition of HIV-1 Nef by a high-affinity binding single-domain antibody.
Lulf, S., Matz, J., Rouyez, M.C., Jarviluoma, A., Saksela, K., Benichou, S., Geyer, M.(2014) Retrovirology 11: 24-24
- PubMed: 24620746 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1742-4690-11-24
- Primary Citation Related Structures:
4ORZ - PubMed Abstract:
The HIV-1 Nef protein is essential for AIDS pathogenesis by its interaction with host cell surface receptors and signaling factors. Despite its critical role as a virulence factor Nef is not targeted by current antiviral strategies. We have determined the crystal structure of the complex formed by a camelid single-domain antibody fragment, termed sdAb19, bound to HIV-1 Nef together with a stabilizing SH3 domain. sdAb19 forms a stoichiometric 1:1 complex with Nef and binds to a conformationally conserved surface at the C-terminus of Nef that overlaps with functionally important interaction sites involved in Nef-induced perturbations of signaling and trafficking pathways. The antibody fragment binds Nef with low nanomolar affinity, which could be attenuated to micromolar affinity range by site-directed mutagenesis of key interaction residues in sdAb19. Fusion of the SH3 domain to sdAb19, termed Neffin, leads to a significantly increased affinity for Nef and formation of a stoichiometric 2:2 Nef-Neffin complex. The 19 kDa Neffin protein inhibits all functions of Nef as CD4 and MHC-I downregulation, association with Pak2, and the increase in virus infectivity and replication. Together, sdAb19 and Neffin thus represent efficient tools for the rational development of antiviral strategies against HIV-1 Nef.
- Center of Advanced European Studies and Research, Group Physical Biochemistry, Bonn, Germany. matthias.geyer@caesar.de.
Organizational Affiliation: