Cooperative autoinhibition and multi-level activation mechanisms of calcineurin
Li, S.J., Ma, L., Wang, J., Lu, C., Wang, J., Wu, J.W., Wang, Z.X.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Serine/threonine-protein phosphatase 2B catalytic subunit beta isoform | 544 | Homo sapiens | Mutation(s): 0 Gene Names: CALNA2, CALNB, CNA2, PPP3CB EC: 3.1.3.16 | 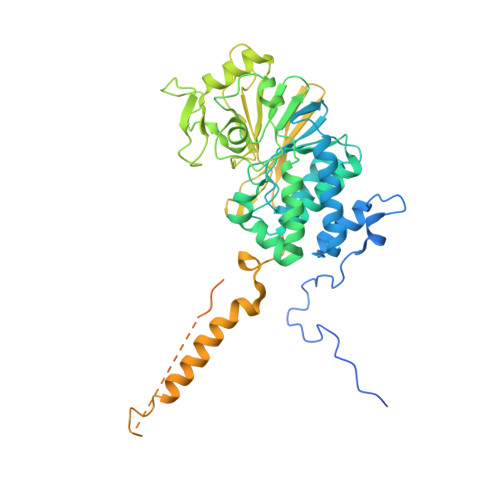 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P16298 GTEx: ENSG00000107758 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P16298 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Calcineurin subunit B type 1 | 170 | Homo sapiens | Mutation(s): 0 Gene Names: CNA2, CNB, PPP3R1 | 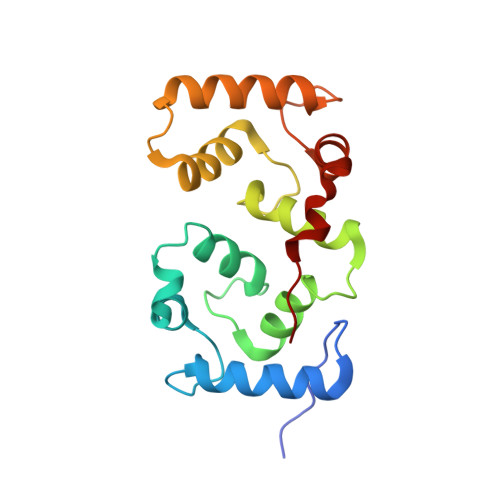 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P63098 GTEx: ENSG00000221823 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P63098 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PO4 Download:Ideal Coordinates CCD File | E [auth A] | PHOSPHATE ION O4 P NBIIXXVUZAFLBC-UHFFFAOYSA-K |  | ||
| ZN Download:Ideal Coordinates CCD File | C [auth A] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| FE Download:Ideal Coordinates CCD File | D [auth A] | FE (III) ION Fe VTLYFUHAOXGGBS-UHFFFAOYSA-N |  | ||
| CA Download:Ideal Coordinates CCD File | F [auth B], G [auth B], H [auth B], I [auth B] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 110.2 | α = 90 |
| b = 110.2 | β = 90 |
| c = 282.488 | γ = 120 |
| Software Name | Purpose |
|---|---|
| HKL-2000 | data collection |
| PHASER | phasing |
| PHENIX | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |