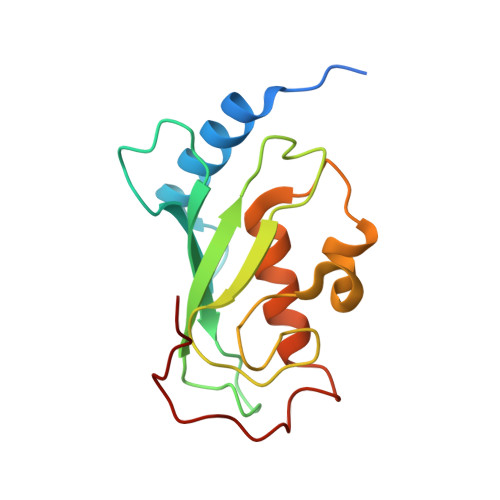
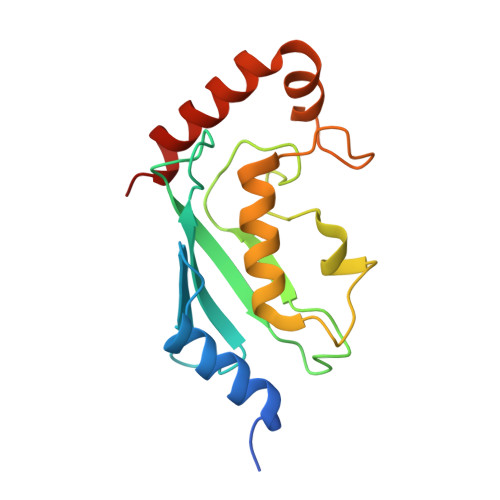
Covalent Inhibition of Ubc13 Affects Ubiquitin Signaling and Reveals Active Site Elements Important for Targeting.
Hodge, C.D., Edwards, R.A., Markin, C.J., McDonald, D., Pulvino, M., Huen, M.S., Zhao, J., Spyracopoulos, L., Hendzel, M.J., Glover, J.N.(2015) ACS Chem Biol 10: 1718-1728
- PubMed: 25909880 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschembio.5b00222
- Primary Citation Related Structures:
4ONL, 4ONM, 4ONN - PubMed Abstract:
Ubc13 is an E2 ubiquitin conjugating enzyme that functions in nuclear DNA damage signaling and cytoplasmic NF-κB signaling. Here, we present the structures of complexes of Ubc13 with two inhibitors, NSC697923 and BAY 11-7082, which inhibit DNA damage and NF-κB signaling in human cells. NSC697923 and BAY 11-7082 both inhibit Ubc13 by covalent adduct formation through a Michael addition at the Ubc13 active site cysteine. The resulting adducts of both compounds exploit a binding groove unique to Ubc13. We developed a Ubc13 mutant which resists NSC697923 inhibition and, using this mutant, we show that the inhibition of cellular DNA damage and NF-κB signaling by NSC697923 is largely due to specific Ubc13 inhibition. We propose that unique structural features near the Ubc13 active site could provide a basis for the rational development and design of specific Ubc13 inhibitors.
- †Department of Biochemistry, University of Alberta, Edmonton, Alberta, Canada T6G 2H7.
Organizational Affiliation: