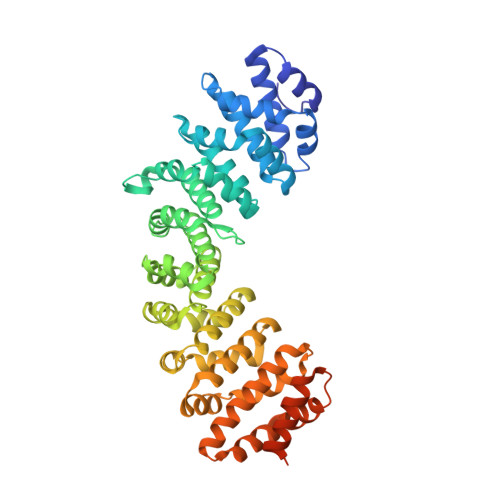
Structural Characterisation of the Nuclear Import Receptor Importin Alpha in Complex with the Bipartite NLS of Prp20
Roman, N., Christie, M., Swarbrick, C.M.D., Kobe, B., Forwood, J.K.(2013) PLoS One 8: e82038-e82038
- PubMed: 24339986 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0082038
- Primary Citation Related Structures:
4OIH - PubMed Abstract:
The translocation of macromolecules into the nucleus is a fundamental eukaryotic process, regulating gene expression, cell division and differentiation, but which is impaired in a range of significant diseases including cancer and viral infection. The import of proteins into the nucleus is generally initiated by a specific, high affinity interaction between nuclear localisation signals (NLSs) and nuclear import receptors in the cytoplasm, and terminated through the disassembly of these complexes in the nucleus. For classical NLSs (cNLSs), this import is mediated by the importin-α (IMPα) adaptor protein, which in turn binds to IMPβ to mediate translocation of nuclear cargo across the nuclear envelope. The interaction and disassembly of import receptor:cargo complexes is reliant on the differential localisation of nucleotide bound Ran across the envelope, maintained in its low affinity, GDP-bound form in the cytoplasm, and its high affinity, GTP-bound form in the nucleus. This in turn is maintained by the differential localisation of Ran regulating proteins, with RanGAP in the cytoplasm maintaining Ran in its GDP-bound form, and RanGEF (Prp20 in yeast) in the nucleus maintaining Ran in its GTP-bound form. Here, we describe the 2.1 Å resolution x-ray crystal structure of IMPα in complex with the NLS of Prp20. We observe 1,091 Å(2) of buried surface area mediated by an extensive array of contacts involving residues on armadillo repeats 2-7, utilising both the major and minor NLS binding sites of IMPα to contact bipartite NLS clusters (17)RAKKMSK(23) and (3)KR(4), respectively. One notable feature of the major site is the insertion of Prp20NLS Ala(18) between the P0 and P1 NLS sites, noted in only a few classical bipartite NLSs. This study provides a detailed account of the binding mechanism enabling Prp20 interaction with the nuclear import receptor, and additional new information for the interaction between IMPα and cargo.
- School of Biomedical Sciences, Charles Sturt University, Wagga Wagga, New South Wales, Australia.
Organizational Affiliation: