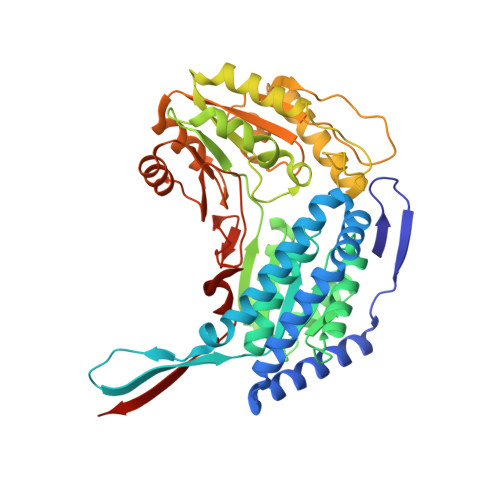
Kinetic and Structural Characterization for Cofactor Preference of Succinic Semialdehyde Dehydrogenase from Streptococcus pyogenes.
Jang, E.H., Park, S.A., Chi, Y.M., Lee, K.S.(2014) Mol Cells 37: 719-726
- PubMed: 25256219
- DOI: https://doi.org/10.14348/molcells.2014.0162
- Primary Citation Related Structures:
4OGD, 4OHT - PubMed Abstract:
The γ-Aminobutyric acid (GABA) that is found in prokaryotic and eukaryotic organisms has been used in various ways as a signaling molecule or a significant component generating metabolic energy under conditions of nutrient limitation or stress, through GABA catabolism. Succinic semialdehyde dehydrogenase (SSADH) catalyzes the oxidation of succinic semialdehyde to succinic acid in the final step of GABA catabolism. Here, we report the catalytic properties and two crystal structures of SSADH from Streptococcus pyogenes (SpSSADH) regarding its cofactor preference. Kinetic analysis showed that SpSSADH prefers NADP(+) over NAD(+) as a hydride acceptor. Moreover, the structures of SpSSADH were determined in an apo-form and in a binary complex with NADP(+) at 1.6 Å and 2.1 Å resolutions, respectively. Both structures of SpSSADH showed dimeric conformation, containing a single cysteine residue in the catalytic loop of each subunit. Further structural analysis and sequence comparison of SpSSADH with other SSADHs revealed that Ser158 and Tyr188 in SpSSADH participate in the stabilization of the 2'-phosphate group of adenine-side ribose in NADP(+). Our results provide structural insights into the cofactor preference of SpSSADH as the gram-positive bacterial SSADH.
- Department of Biosystems and Biotechnology, College of Life Sciences and Biotechnology, Korea University, Seoul 136-713, Korea.
Organizational Affiliation: