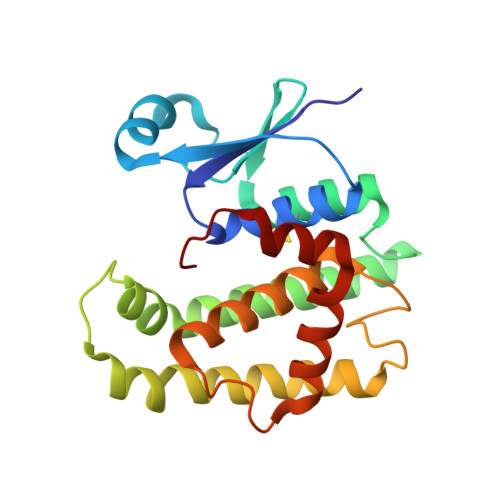
Structure of glutathione S-transferase 1 from the major human hookworm parasite Necator americanus (Na-GST-1) in complex with glutathione.
Asojo, O.A., Ceccarelli, C.(2014) Acta Crystallogr F Struct Biol Commun 70: 1162-1166
- PubMed: 25195885 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X1401646X
- Primary Citation Related Structures:
4OFM, 4OFN, 4OFT - PubMed Abstract:
Glutathione S-transferase 1 from Necator americanus (Na-GST-1) is a vaccine candidate for hookworm infection that has a high affinity for heme and metal porphyrins. As part of attempts to clarify the mechanism of heme detoxification by hookworm GSTs, co-crystallization and soaking studies of Na-GST-1 with the heme-like molecules protoporphyrin IX disodium salt, hematin and zinc protoporphyrin were undertaken. While these studies did not yield the structure of the complex of Na-GST-1 with any of these molecules, co-crystallization experiments resulted in the first structures of the complex of Na-GST-1 with the substrate glutathione. The structures of the complex of Na-GST-1 with glutathione were solved from pathological crystalline aggregates comprising more than one crystal form. These first structures of the complex of Na-GST-1 with the substrate glutathione were solved by molecular replacement from data collected with a sealed-tube home source using the previously reported apo structure as the search model.
- National School of Tropical Medicine, Baylor College of Medicine, Houston, TX 77030, USA.
Organizational Affiliation: