Discovery of 5,6,7,8-tetrahydropyrido[3,4-d]pyrimidine inhibitors of Erk2.
Blake, J.F., Gaudino, J.J., De Meese, J., Mohr, P., Chicarelli, M., Tian, H., Garrey, R., Thomas, A., Siedem, C.S., Welch, M.B., Kolakowski, G., Kaus, R., Burkard, M., Martinson, M., Chen, H., Dean, B., Dudley, D.A., Gould, S.E., Pacheco, P., Shahidi-Latham, S., Wang, W., West, K., Yin, J., Moffat, J., Schwarz, J.B.(2014) Bioorg Med Chem Lett 24: 2635-2639
- PubMed: 24813737 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2014.04.068
- Primary Citation Related Structures:
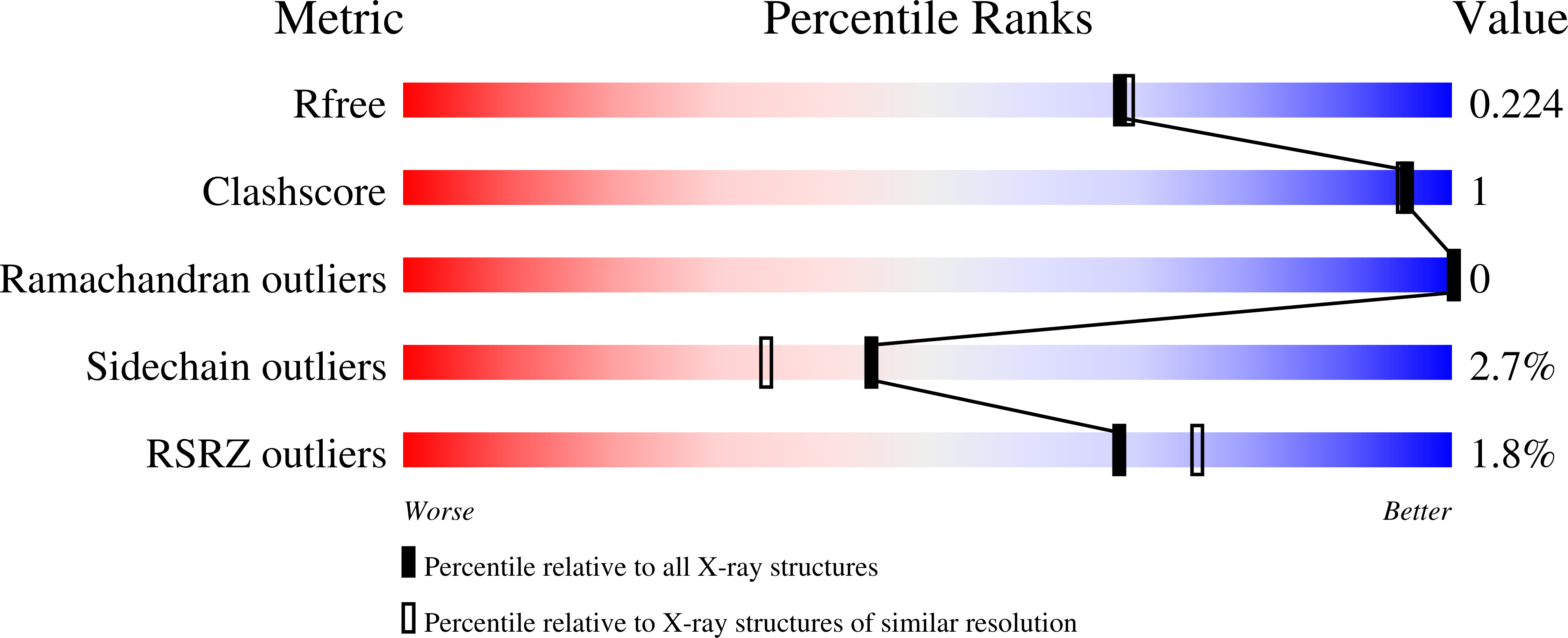
4O6E - PubMed Abstract:
The discovery and optimization of a series of tetrahydropyridopyrimidine based extracellular signal-regulated kinase (Erks) inhibitors discovered via HTS and structure based drug design is reported. The compounds demonstrate potent and selective inhibition of Erk2 and knockdown of phospho-RSK levels in HepG2 cells and tumor xenografts.
- Array BioPharma Inc, Boulder, CO 80301, USA. Electronic address: jblake@arraybiopharma.com.
Organizational Affiliation: