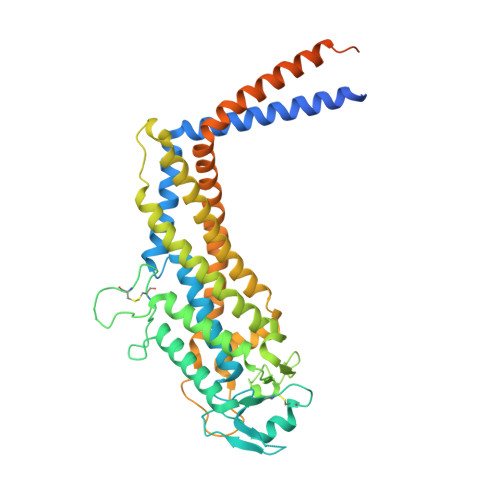
The three-dimensional structure of the extracellular adhesion domain of the sialic acid-binding adhesin SabA from Helicobacter pylori
Pang, S.S., Nguyen, S.T.S., Perry, A.J., Day, C.J., Panjikar, S., Tiralongo, J., Whisstock, J.C., Kwok, T.(2013) J Biological Chem 289: 6332-6340
- PubMed: 24375407 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.513135
- Primary Citation Related Structures:
4O5J - PubMed Abstract:
The gastric pathogen Helicobacter pylori is a major cause of acute chronic gastritis and the development of stomach and duodenal ulcers. Chronic infection furthermore predisposes to the development of gastric cancer. Crucial to H. pylori survival within the hostile environment of the digestive system are the adhesins SabA and BabA; these molecules belong to the same protein family and permit the bacteria to bind tightly to sugar moieties Lewis(B) and sialyl-Lewis(X), respectively, on the surface of epithelial cells lining the stomach and duodenum. To date, no representative SabA/BabA structure has been determined, hampering the development of strategies to eliminate persistent H. pylori infections that fail to respond to conventional therapy. Here, using x-ray crystallography, we show that the soluble extracellular adhesin domain of SabA shares distant similarity to the tetratricopeptide repeat fold family. The molecule broadly resembles a golf putter in shape, with the head region featuring a large cavity surrounded by loops that vary in sequence between different H. pylori strains. The N-terminal and C-terminal helices protrude at right angles from the head domain and together form a shaft that connects to a predicted outer membrane protein-like β-barrel trans-membrane domain. Using surface plasmon resonance, we were able to detect binding of the SabA adhesin domain to sialyl-Lewis(X) and Lewis(X) but not to Lewis(A), Lewis(B), or Lewis(Y). Substitution of the highly conserved glutamine residue 159 in the predicted ligand-binding pocket abrogates the binding of the SabA adhesin domain to sialyl-Lewis(X) and Lewis(X). Taken together, these data suggest that the adhesin domain of SabA is sufficient in isolation for specific ligand binding.
- Department of Biochemistry and Molecular Biology, Monash University, Clayton Campus, Melbourne, Victoria 3800 Australia.
Organizational Affiliation: