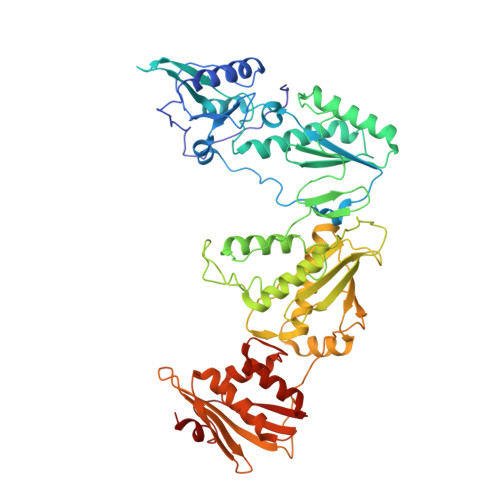
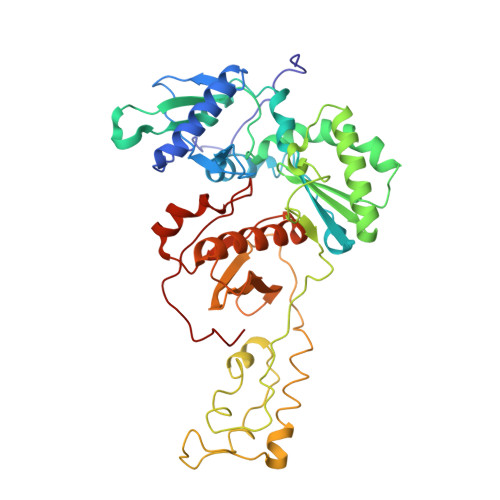
A mechanistic and structural investigation of modified derivatives of the diaryltriazine class of NNRTIs targeting HIV-1 reverse transcriptase.
Mislak, A.C., Frey, K.M., Bollini, M., Jorgensen, W.L., Anderson, K.S.(2014) Biochim Biophys Acta 1840: 2203-2211
- PubMed: 24726448 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbagen.2014.04.001
- Primary Citation Related Structures:
4O44, 4O4G - PubMed Abstract:
Non-nucleoside reverse transcriptase inhibitors (NNRTIs) are vital in treating HIV-1 infection by inhibiting reverse transcriptase (RT). Drug toxicity and resistance drive the need for effective new inhibitors with improved physiochemical properties and potent antiviral activity. Computer-aided and structure-based drug design have guided the addition of solubilizing substituents to the diaryltriazine scaffold. These derivatives have markedly improved solubility and maintain low nanomolar antiviral activity against RT. The molecular and structural basis of inhibition for this series was determined to facilitate future inhibitor development with improved pharmacological profiles. The molecular mechanism of inhibition was investigated using transient-state kinetic analysis. Crystal structures of RT in complex with each inhibitor were obtained to investigate the structural basis of inhibition. The diaryltriazine and its morpholine derivative have RT inhibition constants of 9±2nM and 14±4nM, respectively. They adopt differential binding modes within the non-nucleoside inhibitor binding pocket to distort the catalytic site geometry and primer grip regions. The novel morpholinopropoxy substituent extends into the RT/solvent interface of the NNIBP. Kinetic and structural analyses show that these inhibitors behave as conventional NNRTIs and inhibit the polymerization step. This study confirms that appending solubilizing substituents on the azine ring of diaryltriazine class of NNRTIs that extend into the RT/solvent interface effectively maintains low nanomolar potency and improves physiochemical properties. The modification of NNRTI scaffolds with solubilizing substituents, which extend into the RT/solvent interface, yields potent antivirals and is an effective strategy for developing novel inhibitors with improved pharmacological properties.
- Department of Pharmacology, Yale University School of Medicine, New Haven, CT 06520-8066, USA.
Organizational Affiliation: