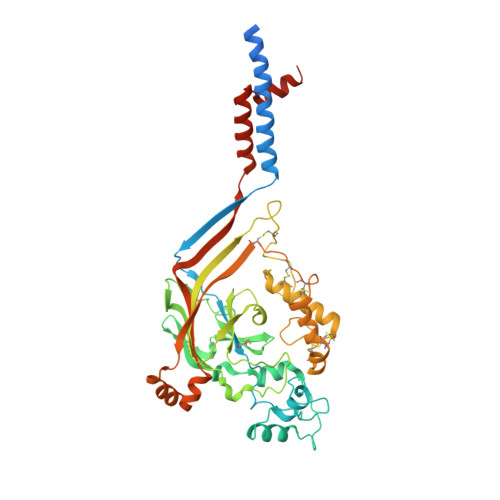
Pore architecture and ion sites in acid-sensing ion channels and P2X receptors.
Gonzales, E.B., Kawate, T., Gouaux, E.(2009) Nature 460: 599-604
- PubMed: 19641589 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature08218
- Primary Citation Related Structures:
3IJ4, 4NYK - PubMed Abstract:
Acid-sensing ion channels are proton-activated, sodium-selective channels composed of three subunits, and are members of the superfamily of epithelial sodium channels, mechanosensitive and FMRF-amide peptide-gated ion channels. These ubiquitous eukaryotic ion channels have essential roles in biological activities as diverse as sodium homeostasis, taste and pain. Despite their crucial roles in biology and their unusual trimeric subunit stoichiometry, there is little knowledge of the structural and chemical principles underlying their ion channel architecture and ion-binding sites. Here we present the structure of a functional acid-sensing ion channel in a desensitized state at 3 A resolution, the location and composition of the approximately 8 A 'thick' desensitization gate, and the trigonal antiprism coordination of caesium ions bound in the extracellular vestibule. Comparison of the acid-sensing ion channel structure with the ATP-gated P2X(4) receptor reveals similarity in pore architecture and aqueous vestibules, suggesting that there are unanticipated yet common structural and mechanistic principles.
- Vollum Institute, Oregon Health and Science University, 3181 Southwest Sam Jackson Park Road, Portland, Oregon 97239, USA.
Organizational Affiliation: