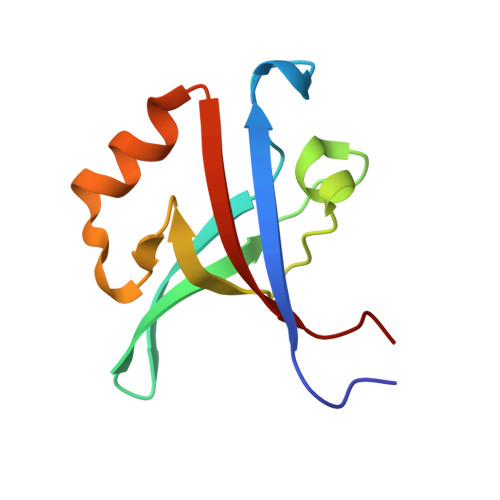
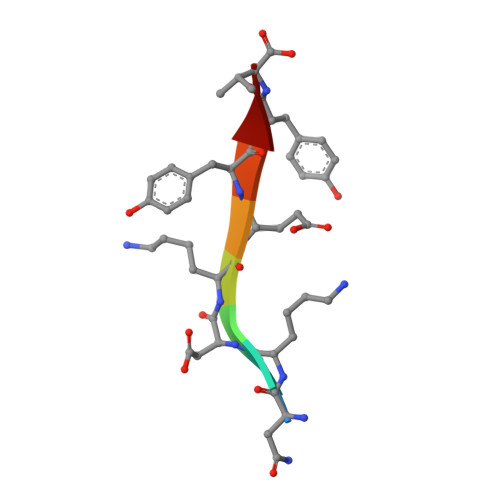
Distinct Roles for Conformational Dynamics in Protein-Ligand Interactions.
Liu, X., Speckhard, D.C., Shepherd, T.R., Sun, Y.J., Hengel, S.R., Yu, L., Fowler, C.A., Gakhar, L., Fuentes, E.J.(2016) Structure 24: 2053-2066
- PubMed: 27998539 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2016.08.019
- Primary Citation Related Structures:
4NXP, 4NXQ, 4NXR - PubMed Abstract:
Conformational dynamics has an established role in enzyme catalysis, but its contribution to ligand binding and specificity is largely unexplored. Here we used the Tiam1 PDZ domain and an engineered variant (QM PDZ) with broadened specificity to investigate the role of structure and conformational dynamics in molecular recognition. Crystal structures of the QM PDZ domain both free and bound to ligands showed structural features central to binding (enthalpy), while nuclear-magnetic-resonance-based methyl relaxation experiments and isothermal titration calorimetry revealed that conformational entropy contributes to affinity. In addition to motions relevant to thermodynamics, slower microsecond to millisecond switching was prevalent in the QM PDZ ligand-binding site consistent with a role in ligand specificity. Our data indicate that conformational dynamics plays distinct and fundamental roles in tuning the affinity (conformational entropy) and specificity (excited-state conformations) of molecular interactions. More broadly, our results have important implications for the evolution, regulation, and design of protein-ligand interactions.
- Department of Biochemistry, University of Iowa, Iowa City, IA 52242-1109, USA; Roy J. and Lucille A. Carver College of Medicine, University of Iowa, Iowa City, IA 52242-1109, USA.
Organizational Affiliation: