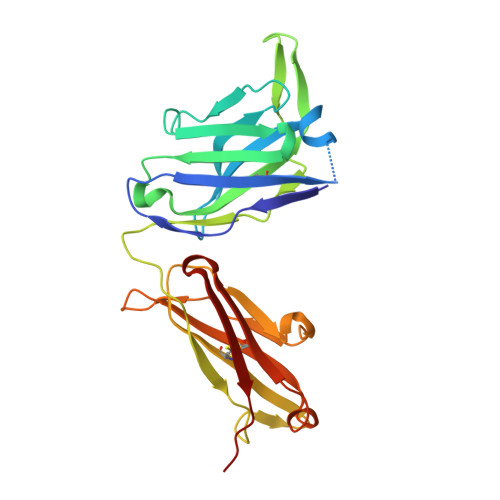
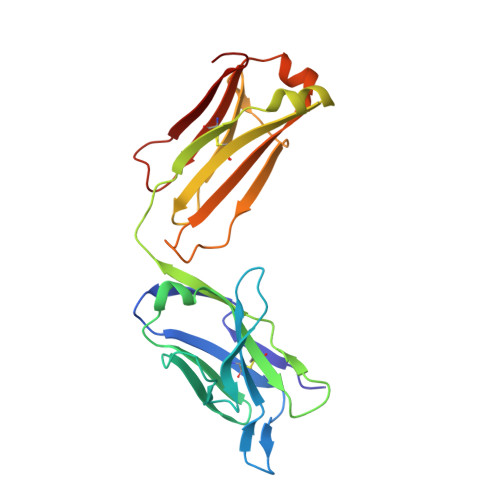
Nucleotide insertions and deletions complement point mutations to massively expand the diversity created by somatic hypermutation of antibodies.
Bowers, P.M., Verdino, P., Wang, Z., da Silva Correia, J., Chhoa, M., Macondray, G., Do, M., Neben, T.Y., Horlick, R.A., Stanfield, R.L., Wilson, I.A., King, D.J.(2014) J Biological Chem 289: 33557-33567
- PubMed: 25320089 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.607176
- Primary Citation Related Structures:
4NWT, 4NWU - PubMed Abstract:
During somatic hypermutation (SHM), deamination of cytidine by activation-induced cytidine deaminase and subsequent DNA repair generates mutations within immunoglobulin V-regions. Nucleotide insertions and deletions (indels) have recently been shown to be critical for the evolution of antibody binding. Affinity maturation of 53 antibodies using in vitro SHM in a non-B cell context was compared with mutation patterns observed for SHM in vivo. The origin and frequency of indels seen during in vitro maturation were similar to that in vivo. Indels are localized to CDRs, and secondary mutations within insertions further optimize antigen binding. Structural determination of an antibody matured in vitro and comparison with human-derived antibodies containing insertions reveal conserved patterns of antibody maturation. These findings indicate that activation-induced cytidine deaminase acting on V-region sequences is sufficient to initiate authentic formation of indels in vitro and in vivo and that point mutations, indel formation, and clonal selection form a robust tripartite system for antibody evolution.
- From Anaptysbio Inc., San Diego, California 92121 and pbowers@anaptysbio.com.
Organizational Affiliation: