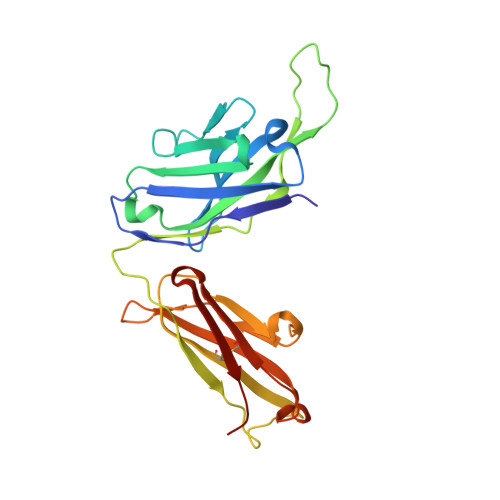
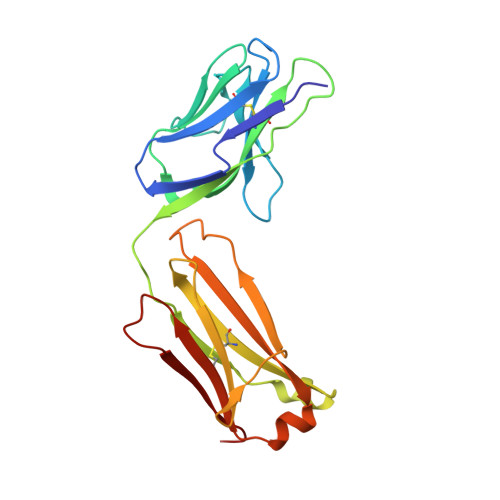
Structural Basis for HIV-1 Neutralization by 2F5-Like Antibodies m66 and m66.6.
Ofek, G., Zirkle, B., Yang, Y., Zhu, Z., McKee, K., Zhang, B., Chuang, G.Y., Georgiev, I.S., O'Dell, S., Doria-Rose, N., Mascola, J.R., Dimitrov, D.S., Kwong, P.D.(2014) J Virol 88: 2426-2441
- PubMed: 24335316
- DOI: https://doi.org/10.1128/JVI.02837-13
- Primary Citation Related Structures:
4NRX, 4NRY, 4NRZ - PubMed Abstract:
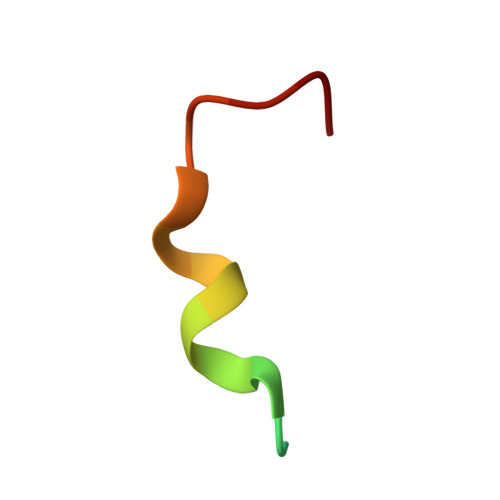
Antibodies m66.6 and 2F5 are the only effective human HIV-1-neutralizing antibodies reported thus far to recognize the N-terminal region of the membrane-proximal external region (MPER) of the gp41 subunit of the HIV-1 envelope glycoprotein. Although 2F5 has been extensively characterized, much less is known about antibody m66.6 or antibody m66, a closely related light-chain variant. Here, we report the crystal structure of m66 in complex with its gp41 epitope, along with unbound structures of m66 and m66.6. We used mutational and binding analyses to decipher antibody elements critical for their recognition of gp41 and determined the molecular basis that underlies their neutralization of HIV-1. When bound by m66, the N-terminal region of the gp41 MPER adopts a conformation comprising a helix, followed by an extended loop. Comparison of gp41-bound m66 to unbound m66.6 identified three light-chain residues of m66.6 that were confirmed through mutagenesis to underlie the greater breadth of m66.6-mediated virus neutralization. Recognition of gp41 by m66 also revealed similarities to antibody 2F5 both in the conformation of crucial epitope residues as well as in the angle of antibody approach. Aromatic residues at the tip of the m66.6 heavy-chain third complementarity-determining region, as in the case of 2F5, were determined to be critical for virus neutralization in a manner that correlated with antibody recognition of the MPER in a lipid context. Antibodies m66, m66.6, and 2F5 thus utilize similar mechanistic elements to recognize a common gp41-MPER epitope and to neutralize HIV-1.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, Maryland, USA.
Organizational Affiliation: