Genome and cancer single nucleotide polymorphisms of the human NEIL1 DNA glycosylase: Activity, structure, and the effect of editing.
Prakash, A., Carroll, B.L., Sweasy, J.B., Wallace, S.S., Doublie, S.(2014) DNA Repair (Amst) 14: 17-26
- PubMed: 24382305 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.dnarep.2013.12.003
- Primary Citation Related Structures:
4NRV, 4NRW - PubMed Abstract:
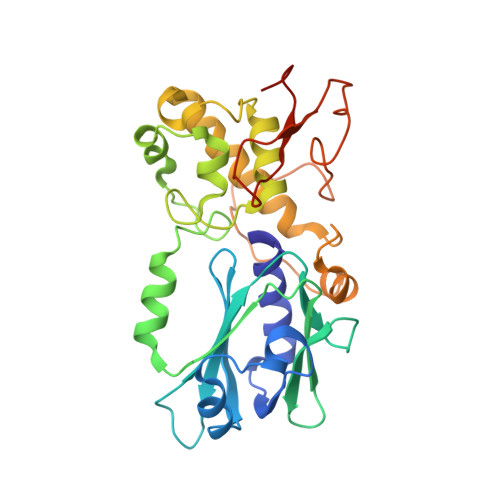
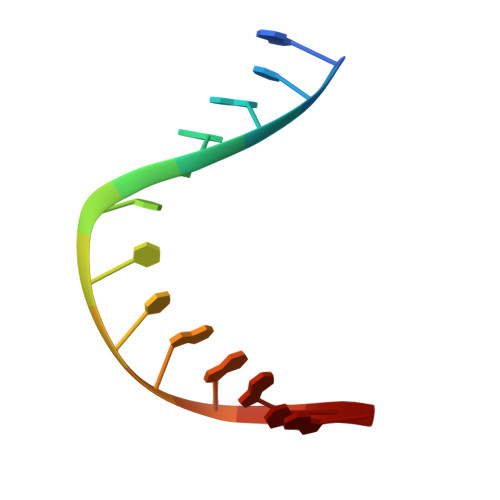
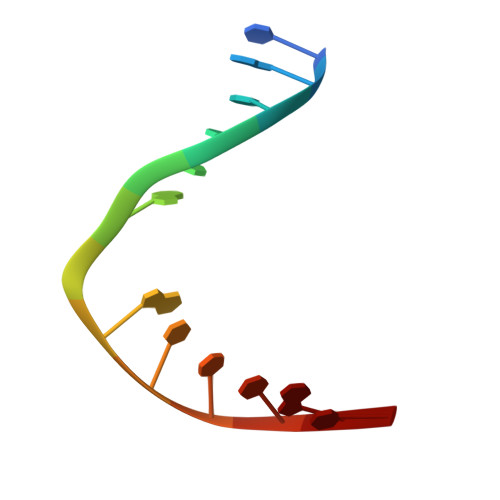
The repair of free-radical oxidative DNA damage is carried out by lesion-specific DNA glycosylases as the first step of the highly conserved base excision repair (BER) pathway. In humans, three orthologs of the prototypical endonuclease VIII (Nei), the Nei-like NEIL1-3 enzymes are involved in the repair of oxidized DNA lesions. In recent years, several genome and cancer single-nucleotide polymorphic variants of the NEIL1 glycosylase have been identified. In this study we characterized four variants of human NEIL1: S82C, G83D, P208S, and ΔE28, and tested their ability to excise pyrimidine-derived lesions such as thymine glycol (Tg), 5-hydroxyuracil (5-OHU), and dihydrouracil (DHU) and the purine-derived guanidinohydantoin (Gh), spiroiminodihydantoin 1 (Sp1), and methylated 2,6-diamino-4-hydroxy-5-formamidopyrimidine (MeFapyG). The P208S variant has near wild-type activity on all substrates tested. The S82C and ΔE28 variants exhibit decreased Tg excision compared to wild-type. G83D displays little to no activity with any of the substrates tested, with the exception of Gh and Sp1. Human NEIL1 is known to undergo editing whereby the lysine at position 242 is recoded into an arginine. The non-edited form of NEIL1 is more efficient at cleaving Tg than the R242 form, but the G83D variant does not cleave Tg regardless of the edited status of NEIL1. The corresponding G86D variant in Mimivirus Nei1 similarly lacks glycosylase activity. A structure of a G86D-DNA complex reveals a rearrangement in the β4/5 loop comprising Leu84, the highly-conserved void-filling residue, thereby providing a structural rationale for the decreased glycosylase activity of the glycine to aspartate variant.
- Department of Microbiology and Molecular Genetics, The Markey Center for Molecular Genetics, University of Vermont, Stafford Hall, 95 Carrigan Drive, Burlington, VT 05405-0068, United States.
Organizational Affiliation: