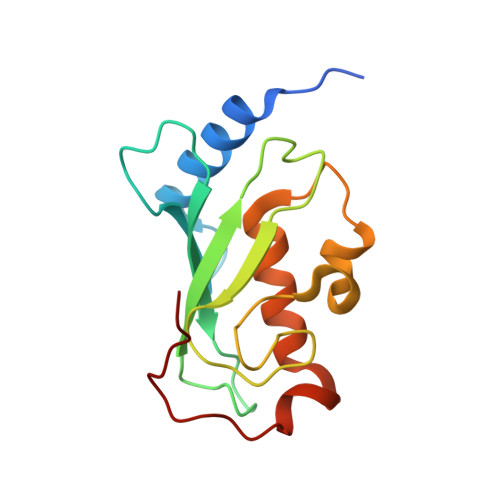
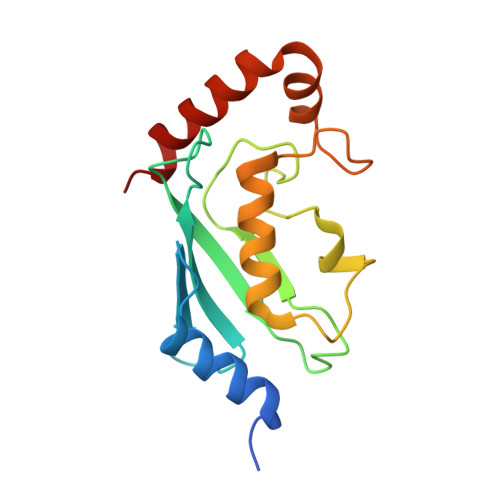
Stochastic gate dynamics regulate the catalytic activity of ubiquitination enzymes.
Rout, M.K., Hodge, C.D., Markin, C.J., Xu, X., Glover, J.N., Xiao, W., Spyracopoulos, L.(2014) J Am Chem Soc 136: 17446-17458
- PubMed: 25423605 Search on PubMed
- DOI: https://doi.org/10.1021/ja505440b
- Primary Citation Related Structures:
4NR3, 4NRG, 4NRI - PubMed Abstract:
Initiation of the DNA damage and innate immune responses is dependent upon the flow of chemical information through coupled protein-protein interaction networks and driven by the synthesis and recognition of Lys 63 linked polyubiquitin (polyUb) chains on adaptor proteins. The central chemical step in Lys 63-linked protein ubiquitination involves the reaction of a specific lysine on a target protein with Ub that is covalently attached as a thioester conjugate to the Ub conjugating enzyme (E2) Ubc13. The active site cysteine of Ubc13, and E2 enzymes in general, is buttressed by a flexible loop. The role of loop dynamics in catalysis was investigated by mutating the central and hinge residues to glycine. The loop dynamics were experimentally characterized through measurement of enzyme kinetics, main chain NMR relaxation, X-ray crystallographic studies, and in vivo studies in yeast. The experimental data were complemented by analysis of MD simulations of the dynamics and kinetics for the loop motion. The results show that fast pico- to nanosecond time scale active site loop fluctuations play a crucial role in regulating the catalytic activity of Ubc13 by functioning as a stochastic active site gate, which is characterized by precisely balanced rates of opening and closing. In vivo functional complementation assays in yeast demonstrate that defects within this regulatory mechanism can have profound biological consequences, given that Ubc13 is the only E2 dedicated to synthesizing Lys 63-linked polyUb chains.
- Department of Biochemistry, University of Alberta , Edmonton, Alberta T6G 2H7, Canada.
Organizational Affiliation: