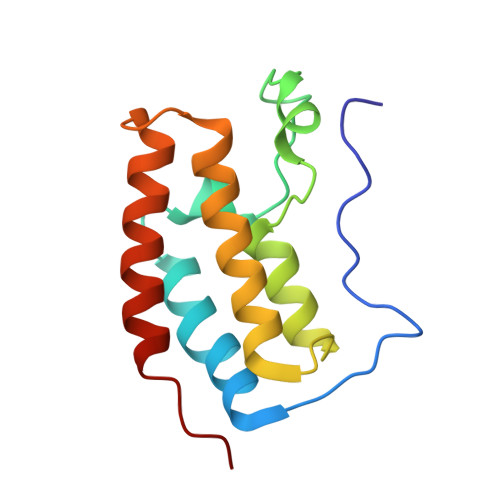
Crystal Structure of the first bromodomain of human BRD4 in complex with a triazolo-phthalazine ligand
Filippakopoulos, P., Picaud, S., Felletar, I., Martin, S., Fedorov, O., von Delft, F., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Knapp, S.To be published.