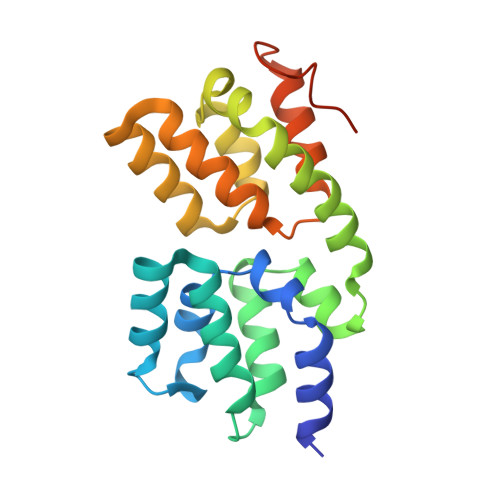
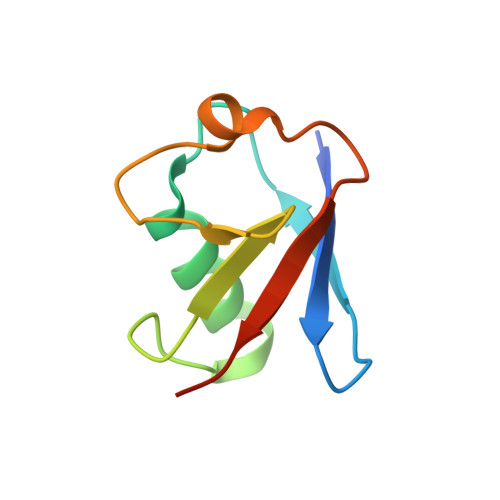
Structural basis for ubiquitin-mediated antiviral signal activation by RIG-I.
Peisley, A., Wu, B., Xu, H., Chen, Z.J., Hur, S.(2014) Nature 509: 110-114
- PubMed: 24590070 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature13140
- Primary Citation Related Structures:
4NQK - PubMed Abstract:
Ubiquitin (Ub) has important roles in a wide range of intracellular signalling pathways. In the conventional view, ubiquitin alters the signalling activity of the target protein through covalent modification, but accumulating evidence points to the emerging role of non-covalent interaction between ubiquitin and the target. In the innate immune signalling pathway of a viral RNA sensor, RIG-I, both covalent and non-covalent interactions with K63-linked ubiquitin chains (K63-Ubn) were shown to occur in its signalling domain, a tandem caspase activation and recruitment domain (hereafter referred to as 2CARD). Non-covalent binding of K63-Ubn to 2CARD induces its tetramer formation, a requirement for downstream signal activation. Here we report the crystal structure of the tetramer of human RIG-I 2CARD bound by three chains of K63-Ub2. 2CARD assembles into a helical tetramer resembling a 'lock-washer', in which the tetrameric surface serves as a signalling platform for recruitment and activation of the downstream signalling molecule, MAVS. Ubiquitin chains are bound along the outer rim of the helical trajectory, bridging adjacent subunits of 2CARD and stabilizing the 2CARD tetramer. The combination of structural and functional analyses reveals that binding avidity dictates the K63-linkage and chain-length specificity of 2CARD, and that covalent ubiquitin conjugation of 2CARD further stabilizes the Ub-2CARD interaction and thus the 2CARD tetramer. Our work provides unique insights into the novel types of ubiquitin-mediated signal-activation mechanism, and previously unexpected synergism between the covalent and non-covalent ubiquitin interaction modes.
- 1] Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, Massachusetts 02115 USA [2] Program in Cellular and Molecular Medicine, Children's Hospital Boston, Boston, Massachusetts 02115, USA.
Organizational Affiliation: