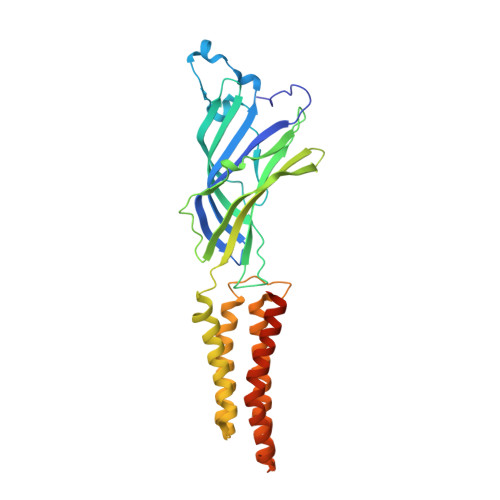
Crystal structures of a pentameric ligand-gated ion channel provide a mechanism for activation.
Sauguet, L., Shahsavar, A., Poitevin, F., Huon, C., Menny, A., Nemecz, A., Haouz, A., Changeux, J.P., Corringer, P.J., Delarue, M.(2014) Proc Natl Acad Sci U S A 111: 966-971
- PubMed: 24367074 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1314997111
- Primary Citation Related Structures:
4NPP, 4NPQ - PubMed Abstract:
Pentameric ligand-gated ion channels mediate fast chemical transmission of nerve signals. The structure of a bacterial proton-gated homolog has been established in its open and locally closed conformations at acidic pH. Here we report its crystal structure at neutral pH, thereby providing the X-ray structures of the two end-points of the gating mechanism in the same pentameric ligand-gated ion channel. The large structural variability in the neutral pH structure observed in the four copies of the pentamer present in the asymmetric unit has been used to analyze the intrinsic fluctuations in this state, which are found to prefigure the transition to the open state. In the extracellular domain (ECD), a marked quaternary change is observed, involving both a twist and a blooming motion, and the pore in the transmembrane domain (TMD) is closed by an upper bend of helix M2 (as in locally closed form) and a kink of helix M1, both helices no longer interacting across adjacent subunits. On the tertiary level, detachment of inner and outer β sheets in the ECD reshapes two essential cavities at the ECD-ECD and ECD-TMD interfaces. The first one is the ligand-binding cavity; the other is close to a known divalent cation binding site in other pentameric ligand-gated ion channels. In addition, a different crystal form reveals that the locally closed and open conformations coexist as discrete ones at acidic pH. These structural results, together with site-directed mutagenesis, physiological recordings, and coarse-grained modeling, have been integrated to propose a model of the gating transition pathway.
- Unité de Dynamique Structurale des Macromolécules, Unité Mixte de Recherche (UMR) 3528 du Centre National de la Recherche Scientifique (CNRS), Institut Pasteur, 75015 Paris, France.
Organizational Affiliation: