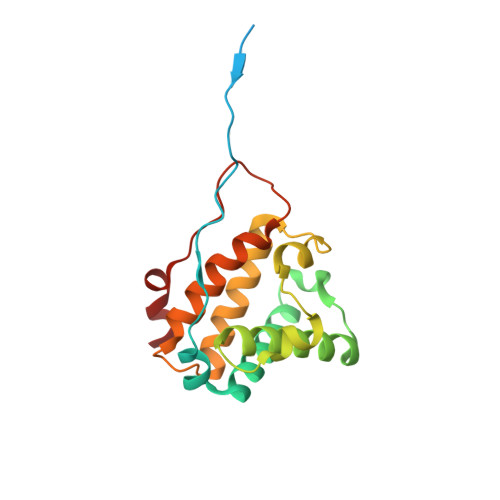
Crystal structure of truncated haemoglobin from an extremely thermophilic and acidophilic bacterium.
Jamil, F., Teh, A.H., Schadich, E., Saito, J.A., Najimudin, N., Alam, M.(2014) J Biochem 156: 97-106
- PubMed: 24733432 Search on PubMed
- DOI: https://doi.org/10.1093/jb/mvu023
- Primary Citation Related Structures:
4NK1, 4NK2 - PubMed Abstract:
A truncated haemoglobin (tHb) has been identified in an acidophilic and thermophilic methanotroph Methylacidiphilium infernorum. Hell's Gate Globin IV (HGbIV) and its related tHbs differ from all other bacterial tHbs due to their distinctively large sequence and polar distal haem pocket residues. Here we report the crystal structure of HGbIV determined at 1.96 Å resolution. The HGbIV structure has the distinctive 2/2 α-helical structure with extensions at both termini. It has a large distal site cavity in the haem pocket surrounded by four polar residues: His70(B9), His71(B10), Ser97(E11) and Trp137(G8). This cavity can bind bulky ligands such as a phosphate ion. Conformational shifts of His71(B10), Leu90(E4) and Leu93(E7) can also provide more space to accommodate larger ligands than the phosphate ion. The entrance/exit of such bulky ligands might be facilitated by positional flexibility in the CD1 loop, E helix and haem-propionate A. Therefore, the large cavity in HGbIV with polar His70(B9) and His71(B10), in contrast to the distal sites of other bacterial tHbs surrounded by non-polar residues, suggests its distinct physiological functions.
- Centre for Chemical Biology, Universiti Sains Malaysia, 10 Persiaran Bukit Jambul, 11900 Bayan Lepas, Penang, Malaysia; School of Biological Sciences, University of Canterbury, Private Bag 4800, Christchurch, New Zealand; Advanced Studies in Genomics, Proteomics and Bioinformatics, University of Hawaii, 2565 McCarthy Mall, Honolulu, HI 96822, USA; School of Biological Sciences, Universiti Sains Malaysia, 11800, Penang, Malaysia; and Department of Microbiology, University of Hawaii, 2538 McCarthy Mall, Honolulu, HI 96822, USA farrukhccb@gmail.com.
Organizational Affiliation: