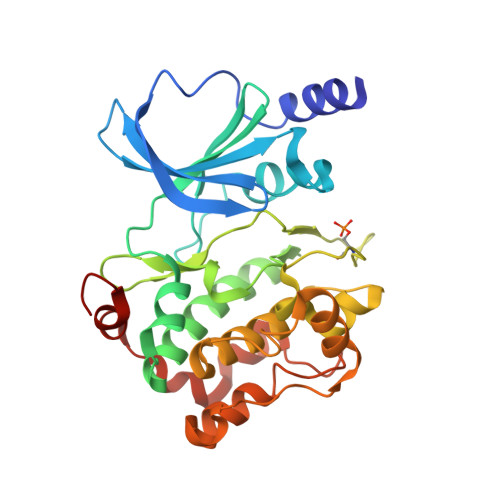
Discovery and the structural basis of a novel p21-activated kinase 4 inhibitor.
Ryu, B.J., Kim, S., Min, B., Kim, K.Y., Lee, J.S., Park, W.J., Lee, H., Kim, S.H., Park, S.(2014) Cancer Lett 349: 45-50
- PubMed: 24704155 Search on PubMed
- DOI: https://doi.org/10.1016/j.canlet.2014.03.024
- Primary Citation Related Structures:
4NJD - PubMed Abstract:
Functional versatility and elevated expression in cancers have endowed p21-activated kinase 4 (PAK4) as one of the first-in-class anti-cancer drug target. In this study, a novel PAK4 inhibitor, KY-04031 (N(2)-(2-(1H-indol-3-yl)ethyl)-N(4)-(1H-indazol-5-yl)-6-methoxy-1,3,5-triazine-2,4-diamine), was discovered using a high-throughput screening. Analysis of the complex crystal structure illustrated that both indole and indazole of KY-04031 are responsible for PAK4 hinge interaction. Moreover, the molecule's triazine core was found to mimic the ribose of the natural ATP substrate. The cell-based anti-cancer potency of KY-04031 was less effective than the pyrroloaminopyrazoles; however, the unique molecular feature of KY-04031 can be exploited in designing new PAK4 inhibitors.
- Laboratory of Translational Therapeutics, Korea Research Institute of Chemical Technology, Yuseong-Gu, Daejeon 305-600, Republic of Korea.
Organizational Affiliation: