Identification of a Small Molecule that Increases Hemoglobin Oxygen Affinity and Reduces SS Erythrocyte Sickling.
Nakagawa, A., Lui, F.E., Wassaf, D., Yefidoff-Freedman, R., Casalena, D., Palmer, M.A., Meadows, J., Mozzarelli, A., Ronda, L., Abdulmalik, O., Bloch, K.D., Safo, M.K., Zapol, W.M.(2014) ACS Chem Biol 9: 2318-2325
- PubMed: 25061917 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/cb500230b
- Primary Citation Related Structures:
4NI0, 4NI1 - PubMed Abstract:
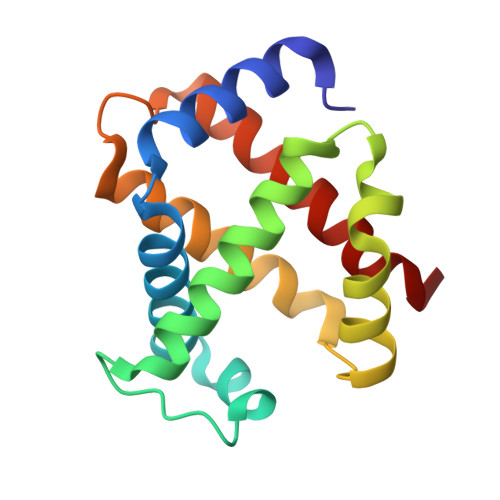
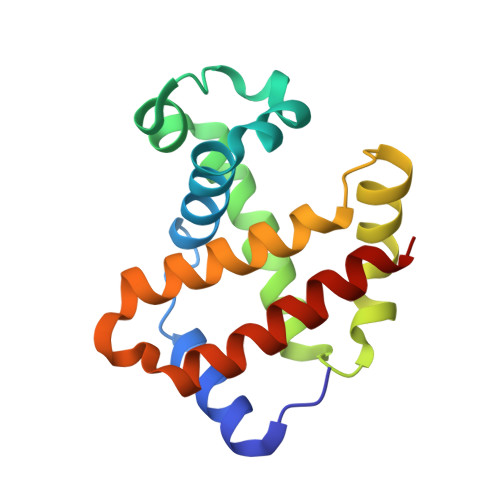
Small molecules that increase the oxygen affinity of human hemoglobin may reduce sickling of red blood cells in patients with sickle cell disease. We screened 38,700 compounds using small molecule microarrays and identified 427 molecules that bind to hemoglobin. We developed a high-throughput assay for evaluating the ability of the 427 small molecules to modulate the oxygen affinity of hemoglobin. We identified a novel allosteric effector of hemoglobin, di(5-(2,3-dihydro-1,4-benzodioxin-2-yl)-4H-1,2,4-triazol-3-yl)disulfide (TD-1). TD-1 induced a greater increase in oxygen affinity of human hemoglobin in solution and in red blood cells than did 5-hydroxymethyl-2-furfural (5-HMF), N-ethylmaleimide (NEM), or diformamidine disulfide. The three-dimensional structure of hemoglobin complexed with TD-1 revealed that monomeric units of TD-1 bound covalently to β-Cys93 and β-Cys112, as well as noncovalently to the central water cavity of the hemoglobin tetramer. The binding of TD-1 to hemoglobin stabilized the relaxed state (R3-state) of hemoglobin. TD-1 increased the oxygen affinity of sickle hemoglobin and inhibited in vitro hypoxia-induced sickling of red blood cells in patients with sickle cell disease without causing hemolysis. Our study indicates that TD-1 represents a novel lead molecule for the treatment of patients with sickle cell disease.
- Anesthesia Center for Critical Care Research, Department of Anesthesia, Critical Care, and Pain Medicine, Massachusetts General Hospital and Harvard Medical School, 55 Fruit Street, Boston, Massachusetts 02114, United States.
Organizational Affiliation: