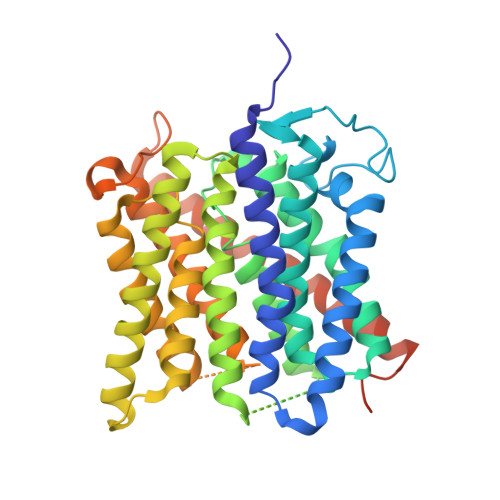
Membrane proteins bind lipids selectively to modulate their structure and function.
Laganowsky, A., Reading, E., Allison, T.M., Ulmschneider, M.B., Degiacomi, M.T., Baldwin, A.J., Robinson, C.V.(2014) Nature 510: 172-175
- PubMed: 24899312
- DOI: https://doi.org/10.1038/nature13419
- Primary Citation of Related Structures:
4NH2 - PubMed Abstract:
Previous studies have established that the folding, structure and function of membrane proteins are influenced by their lipid environments and that lipids can bind to specific sites, for example, in potassium channels. Fundamental questions remain however regarding the extent of membrane protein selectivity towards lipids. Here we report a mass spectrometry approach designed to determine the selectivity of lipid binding to membrane protein complexes. We investigate the mechanosensitive channel of large conductance (MscL) from Mycobacterium tuberculosis and aquaporin Z (AqpZ) and the ammonia channel (AmtB) from Escherichia coli, using ion mobility mass spectrometry (IM-MS), which reports gas-phase collision cross-sections. We demonstrate that folded conformations of membrane protein complexes can exist in the gas phase. By resolving lipid-bound states, we then rank bound lipids on the basis of their ability to resist gas phase unfolding and thereby stabilize membrane protein structure. Lipids bind non-selectively and with high avidity to MscL, all imparting comparable stability; however, the highest-ranking lipid is phosphatidylinositol phosphate, in line with its proposed functional role in mechanosensation. AqpZ is also stabilized by many lipids, with cardiolipin imparting the most significant resistance to unfolding. Subsequently, through functional assays we show that cardiolipin modulates AqpZ function. Similar experiments identify AmtB as being highly selective for phosphatidylglycerol, prompting us to obtain an X-ray structure in this lipid membrane-like environment. The 2.3 Å resolution structure, when compared with others obtained without lipid bound, reveals distinct conformational changes that re-position AmtB residues to interact with the lipid bilayer. Our results demonstrate that resistance to unfolding correlates with specific lipid-binding events, enabling a distinction to be made between lipids that merely bind from those that modulate membrane protein structure and/or function. We anticipate that these findings will be important not only for defining the selectivity of membrane proteins towards lipids, but also for understanding the role of lipids in modulating protein function or drug binding.
- Department of Chemistry, University of Oxford, South Parks Road, Oxford, OX1 5QY, UK.
Organizational Affiliation: