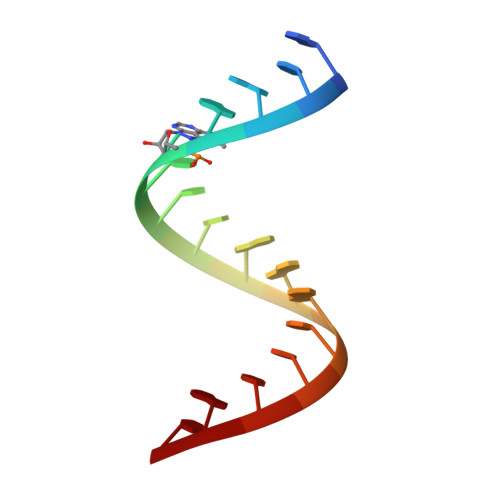
Click Modification of RNA at Adenosine: Structure and Reactivity of 7-Ethynyl- and 7-Triazolyl-8-aza-7-deazaadenosine in RNA.
Phelps, K.J., Ibarra-Soza, J.M., Tran, K., Fisher, A.J., Beal, P.A.(2014) ACS Chem Biol 9: 1780-1787
- PubMed: 24896732 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/cb500270x
- Primary Citation Related Structures:
4NFO, 4NFP, 4NFQ - PubMed Abstract:
Ribonucleoside analogues bearing terminal alkynes, including 7-ethynyl-8-aza-7-deazaadenosine (7-EAA), are useful for RNA modification applications. However, although alkyne- and triazole-bearing ribonucleosides are in widespread use, very little information is available on the impact of these modifications on RNA structure. By solving crystal structures for RNA duplexes containing these analogues, we show that, like adenosine, 7-EAA and a triazole derived from 7-EAA base pair with uridine and are well-accommodated within an A-form helix. We show that copper-catalyzed azide/alkyne cycloaddition (CuAAC) reactions with 7-EAA are sensitive to the RNA secondary structure context, with single-stranded sites reacting faster than duplex sites. 7-EAA and its triazole products are recognized in RNA template strands as adenosine by avian myoblastosis virus reverse transcriptase. In addition, 7-EAA in RNA is a substrate for an active site mutant of the RNA editing adenosine deaminase, ADAR2. These studies extend our understanding of the impact of these novel nucleobase analogues and set the stage for their use in probing RNA structure and metabolism.
- Department of Chemistry, University of California, Davis , One Shields Avenue, Davis, California 95616, United States.
Organizational Affiliation: