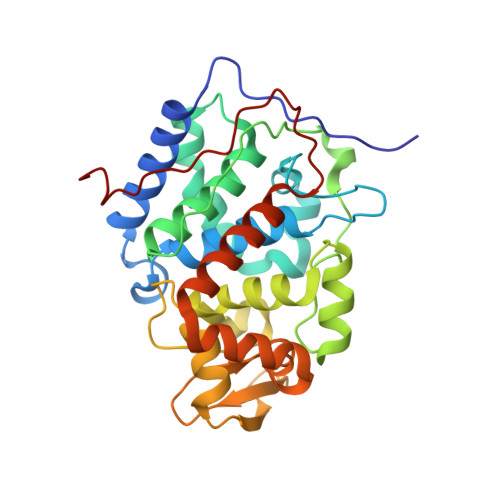
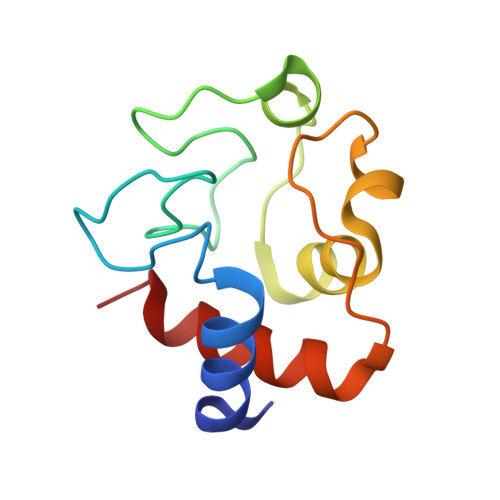
Engineering specificity in a dynamic protein complex with a single conserved mutation.
Bashir, Q., Meulenbroek, E.M., Pannu, N.S., Ubbink, M.(2014) FEBS J 281: 4892-4905
- PubMed: 25180929 Search on PubMed
- DOI: https://doi.org/10.1111/febs.13028
- Primary Citation Related Structures:
4NFG - PubMed Abstract:
It has been demonstrated that the complex of yeast cytochrome c (Cc) and cytochrome c peroxidase (CcP) exists as a delicate equilibrium of a specific, active state and the non-specific, dynamic encounter state. An ortholog of yeast Cc, horse Cc, binds CcP but forms a much more dynamic complex, as demonstrated by NMR spectroscopy. A single conservative mutation of lysine 13 to arginine reduces the dynamics and enhances the specificity. The crystal structure of the stereospecific complex resembles the yeast Cc-CcP complex. In contrast, the K13A mutation increases the dynamic nature of the complex with CcP, showing that specificity in a redox protein complex can depend on the interactions of a single side chain in the binding interface.
- Gorlaeus Laboratories, Leiden Institute of Chemistry, Leiden University, The Netherlands.
Organizational Affiliation: