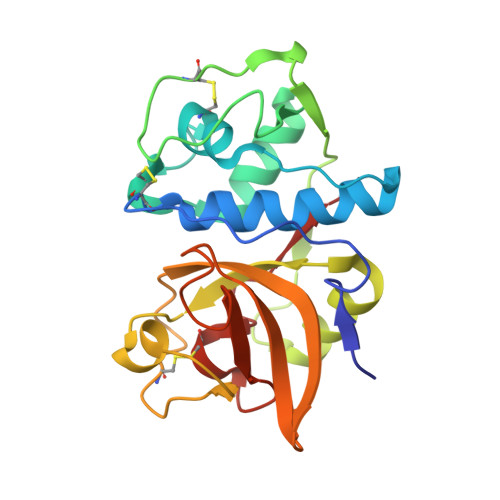
Structural basis of collagen fiber degradation by cathepsin K.
Aguda, A.H., Panwar, P., Du, X., Nguyen, N.T., Brayer, G.D., Bromme, D.(2014) Proc Natl Acad Sci U S A 111: 17474-17479
- PubMed: 25422423 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1414126111
- Primary Citation Related Structures:
4N79, 4N8W - PubMed Abstract:
Cathepsin K is the major collagenolytic protease in bone that facilitates physiological as well as pathological bone degradation. Despite its key role in bone remodeling and for being a highly sought-after drug target for the treatment of osteoporosis, the mechanism of collagen fiber degradation by cathepsin K remained elusive. Here, we report the structure of a collagenolytically active cathepsin K protein dimer. Cathepsin K is organized into elongated C-shaped protease dimers that reveal a putative collagen-binding interface aided by glycosaminoglycans. Molecular modeling of collagen binding to the dimer indicates the participation of nonactive site amino acid residues, Q21 and Q92, in collagen unfolding. Mutations at these sites as well as perturbation of the dimer protein-protein interface completely inhibit cathepsin-K-mediated fiber degradation without affecting the hydrolysis of gelatin or synthetic peptide. Using scanning electron microscopy, we demonstrate the specific binding of cathepsin K at the edge of the fibrillar gap region of collagen fibers, which suggest initial cleavage events at the N- and C-terminal ends of tropocollagen molecules. Edman degradation analysis of collagen fiber degradation products revealed those initial cleavage sites. We propose that one cathepsin K molecule binds to collagen-bound glycosaminoglycans at the gap region and recruits a second protease molecule that provides an unfolding and cleavage mechanism for triple helical collagen. Removal of collagen-associated glycosaminoglycans prevents cathepsin K binding and subsequently fiber hydrolysis. Cathepsin K dimer and glycosaminoglycan binding sites represent novel targeting sites for the development of nonactive site-directed second-generation inhibitors of this important drug target.
- Department of Oral Biological and Medical Sciences, Faculty of Dentistry, Department of Biochemistry and Molecular Biology, Faculty of Medicine, and.
Organizational Affiliation: