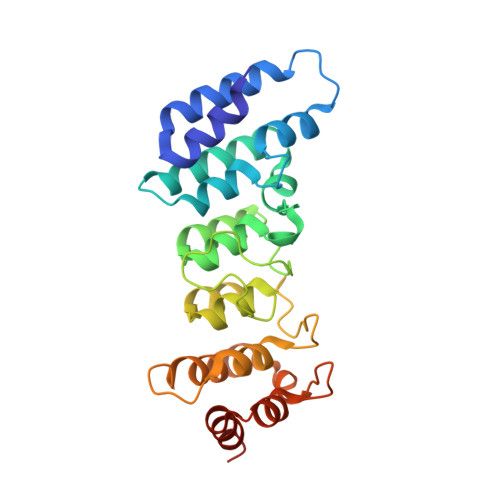
Crystal structure of the N-terminal ankyrin repeat domain of TRPV3 reveals unique conformation of finger 3 loop critical for channel function
Shi, D.J., Ye, S., Cao, X., Zhang, R., Wang, K.(2013) Protein Cell 4: 942-950
- PubMed: 24248473
- DOI: https://doi.org/10.1007/s13238-013-3091-0
- Primary Citation of Related Structures:
4N5Q - PubMed Abstract:
In all six members of TRPV channel subfamily, there is an ankyrin repeat domain (ARD) in their intracellular N-termini. Ankyrin (ANK) repeat, a common motif with typically 33 residues in each repeat, is primarily involved in protein-protein interactions. Despite the sequence similarity among the ARDs of TRPV channels, the structure of TRPV3-ARD, however, remains unknown. Here, we report the crystal structure of TRPV3-ARD solved at 1.95 Å resolution, which reveals six-ankyrin repeats. While overall structure of TRPV3-ARD is similar to ARDs from other members of TRPV subfamily; it, however, features a noticeable finger 3 loop that bends over and is stabilized by a network of hydrogen bonds and hydrophobic packing, instead of being flexible as seen in known TRPV-ARD structures. Electrophysiological recordings demonstrated that mutating key residues R225, R226, Q255, and F249 of finger 3 loop altered the channel activities and pharmacology. Taken all together, our findings show that TRPV3-ARD with characteristic finger 3 loop likely plays an important role in channel function and pharmacology.
- Department of Neurobiology, Neuroscience Research Institute, Peking University Health Science Center, Beijing, 100191, China.
Organizational Affiliation: