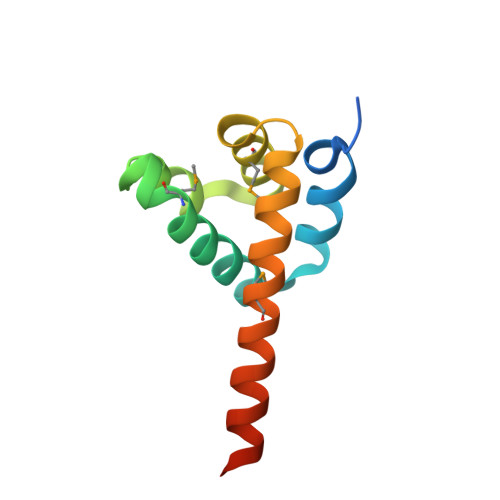
Structures of the NLRP14 pyrin domain reveal a conformational switch mechanism regulating its molecular interactions.
Eibl, C., Hessenberger, M., Wenger, J., Brandstetter, H.(2014) Acta Crystallogr D Biol Crystallogr 70: 2007-2018
- PubMed: 25004977 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004714010311
- Primary Citation Related Structures:
4N1J, 4N1K, 4N1L - PubMed Abstract:
The cytosolic tripartite NLR receptors serve as important signalling platforms in innate immunity. While the C-terminal domains act as sensor and activation modules, the N-terminal death-like domain, e.g. the CARD or pyrin domain, is thought to recruit downstream effector molecules by homotypic interactions. Such homotypic complexes have been determined for all members of the death-domain superfamily except for pyrin domains. Here, crystal structures of human NLRP14 pyrin-domain variants are reported. The wild-type protein as well as the clinical D86V mutant reveal an unexpected rearrangement of the C-terminal helix α6, resulting in an extended α5/6 stem-helix. This reordering mediates a novel symmetric pyrin-domain dimerization mode. The conformational switching is controlled by a charge-relay system with a drastic impact on protein stability. How the identified charge relay allows classification of NLRP receptors with respect to distinct recruitment mechanisms is discussed.
- Department of Molecular Biology, University of Salzburg, Billrothstrasse 11, 5020 Salzburg, Austria.
Organizational Affiliation: