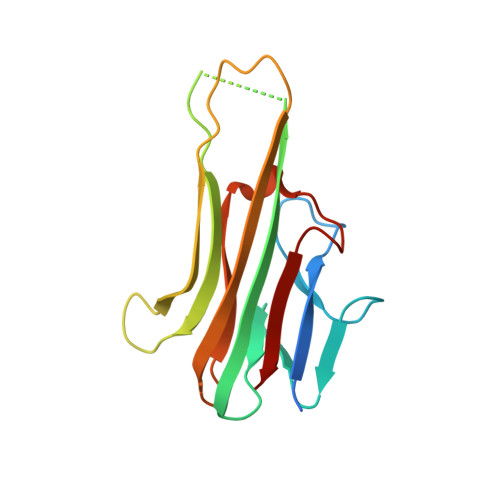
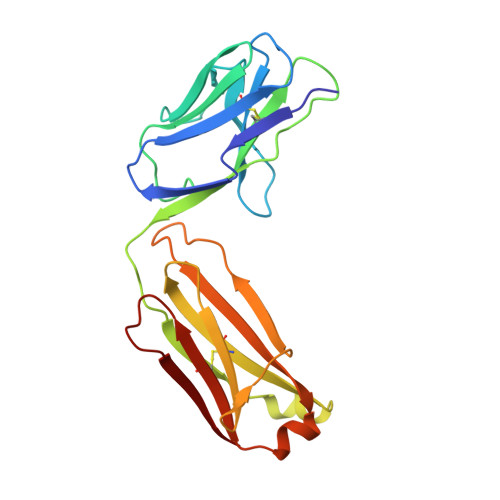
Dimerization of LT beta R by LT alpha 1 beta 2 is necessary and sufficient for signal transduction.
Sudhamsu, J., Yin, J., Chiang, E.Y., Starovasnik, M.A., Grogan, J.L., Hymowitz, S.G.(2013) Proc Natl Acad Sci U S A 110: 19896-19901
- PubMed: 24248355 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1310838110
- Primary Citation Related Structures:
4MXV, 4MXW - PubMed Abstract:
Homotrimeric TNF superfamily ligands signal by inducing trimers of their cognate receptors. As a biologically active heterotrimer, Lymphotoxin(LT)α1β2 is unique in the TNF superfamily. How the three unique potential receptor-binding interfaces in LTα1β2 trigger signaling via LTβ Receptor (LTβR) resulting in lymphoid organogenesis and propagation of inflammatory signals is poorly understood. Here we show that LTα1β2 possesses two binding sites for LTβR with distinct affinities and that dimerization of LTβR by LTα1β2 is necessary and sufficient for signal transduction. The crystal structure of a complex formed by LTα1β2, LTβR, and the fab fragment of an antibody that blocks LTβR activation reveals the lower affinity receptor-binding site. Mutations targeting each potential receptor-binding site in an engineered single-chain variant of LTα1β2 reveal the high-affinity site. NF-κB reporter assays further validate that disruption of receptor interactions at either site is sufficient to prevent signaling via LTβR.
- Departments of Structural Biology and Immunology, Genentech, Inc., South San Francisco, CA 94080.
Organizational Affiliation: