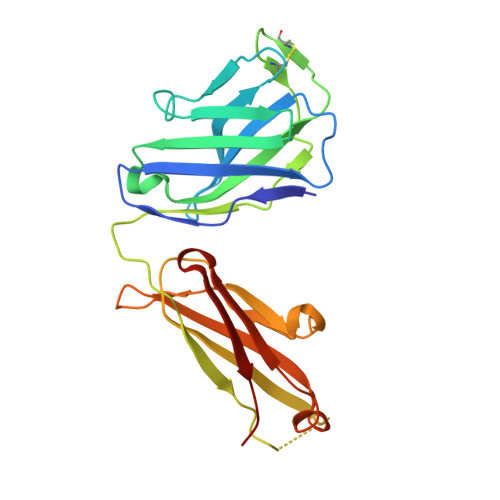
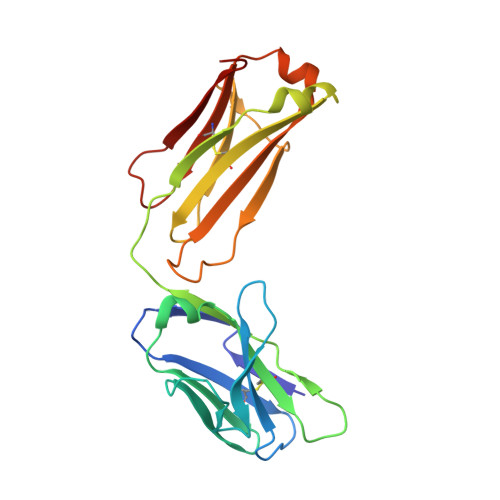
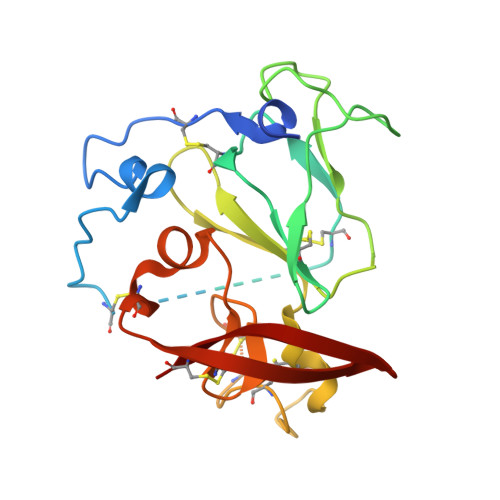
Hepatitis C virus e2 envelope glycoprotein core structure.
Kong, L., Giang, E., Nieusma, T., Kadam, R.U., Cogburn, K.E., Hua, Y., Dai, X., Stanfield, R.L., Burton, D.R., Ward, A.B., Wilson, I.A., Law, M.(2013) Science 342: 1090-1094
- PubMed: 24288331 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1243876
- Primary Citation Related Structures:
4MWF - PubMed Abstract:
Hepatitis C virus (HCV), a Hepacivirus, is a major cause of viral hepatitis, liver cirrhosis, and hepatocellular carcinoma. HCV envelope glycoproteins E1 and E2 mediate fusion and entry into host cells and are the primary targets of the humoral immune response. The crystal structure of the E2 core bound to broadly neutralizing antibody AR3C at 2.65 angstroms reveals a compact architecture composed of a central immunoglobulin-fold β sandwich flanked by two additional protein layers. The CD81 receptor binding site was identified by electron microscopy and site-directed mutagenesis and overlaps with the AR3C epitope. The x-ray and electron microscopy E2 structures differ markedly from predictions of an extended, three-domain, class II fusion protein fold and therefore provide valuable information for HCV drug and vaccine design.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: