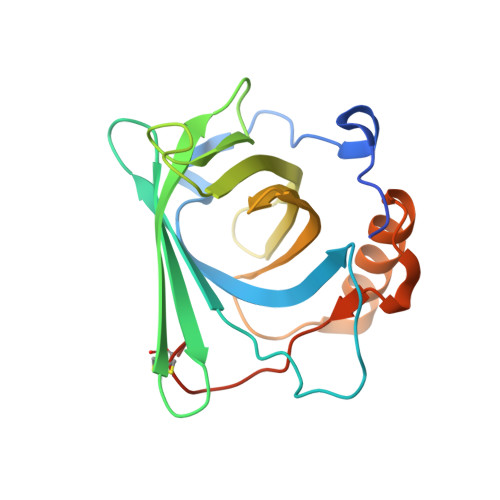
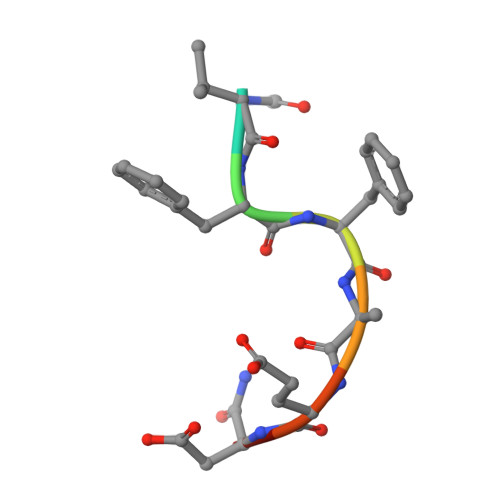
High-affinity Anticalins with aggregation-blocking activity directed against the Alzheimer beta-amyloid peptide.
Rauth, S., Hinz, D., Borger, M., Uhrig, M., Mayhaus, M., Riemenschneider, M., Skerra, A.(2016) Biochem J 473: 1563-1578
- PubMed: 27029347 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BCJ20160114
- Primary Citation Related Structures:
4MVI, 4MVK, 4MVL - PubMed Abstract:
Amyloid beta (Aβ) peptides, in particular Aβ42 and Aβ40, exert neurotoxic effects and their overproduction leads to amyloid deposits in the brain, thus constituting an important biomolecular target for treatments of Alzheimer's disease (AD). We describe the engineering of cognate Anticalins as a novel type of neutralizing protein reagent based on the human lipocalin scaffold. Phage display selection from a genetic random library comprising variants of the human lipocalin 2 (Lcn2) with mutations targeted at 20 exposed amino acid positions in the four loops that form the natural binding site was performed using both recombinant and synthetic target peptides and resulted in three different Anticalins. Biochemical characterization of the purified proteins produced by periplasmic secretion in Escherichia coli revealed high folding stability in a monomeric state, with Tm values ranging from 53.4°C to 74.5°C, as well as high affinities for Aβ40, between 95 pM and 563 pM, as measured by real-time surface plasmon resonance analysis. The central linear VFFAED epitope within the Aβ sequence was mapped using a synthetic peptide array on membranes and was shared by all three Anticalins, despite up to 13 mutual amino acid differences in their binding sites. All Anticalins had the ability-with varying extent-to inhibit Aβ aggregation in vitro according to the thioflavin-T fluorescence assay and, furthermore, they abolished Aβ42-mediated toxicity in neuronal cell culture. Thus, these Anticalins provide not only useful protein reagents to study the molecular pathology of AD but they also show potential as alternative drug candidates compared with antibodies.
- Munich Center for Integrated Protein Science (CIPS-M) and Lehrstuhl für Biologische Chemie, Technische Universität München, 85354 Freising (Weihenstephan), Germany.
Organizational Affiliation: