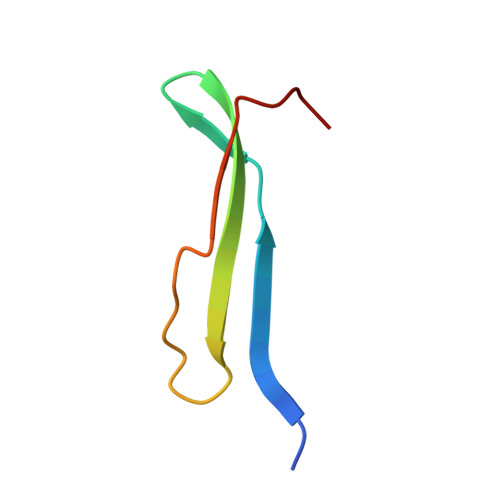
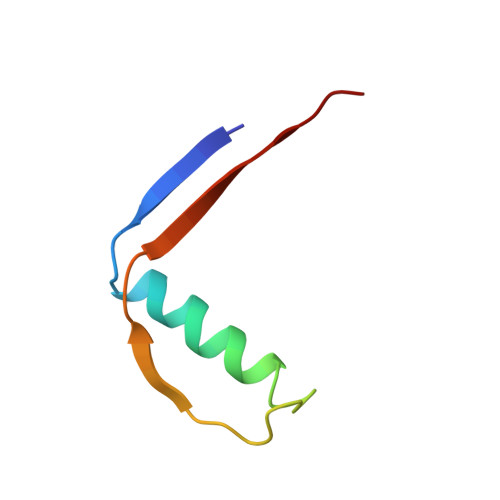
Structure of monellin refined to 2.3 a resolution in the orthorhombic crystal form.
Bujacz, G., Miller, M., Harrison, R., Thanki, N., Gilliland, G.L., Ogata, C.M., Kim, S.H., Wlodawer, A.(1997) Acta Crystallogr D Biol Crystallogr 53: 713-719
- PubMed: 15299859 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444997006860
- Primary Citation Related Structures:
4MON - PubMed Abstract:
The structure of orthorhombic crystals of monellin, a sweet protein extracted from African serendipity berries, has been solved by molecular replacement and refined to 2.3 A resolution. The final R factor was 0.150 for a model with excellent geometry. A monellin molecule consists of two peptides that are non-covalently bound, with chain A composed of three beta-strands interconnected by loop regions and chain B composed of two beta-strands interconnected by an alpha-helix. The N terminus of chain A is in close proximity to the C terminus of chain B. The two molecules in the asymmetric unit are related by a non-crystallographic twofold axis and form a dimer, similar to those previously observed in other crystal forms of both natural and single-chain monellin. The r.m.s, deviation between the Calpha atoms in the two independent molecules is 0.60 A, while the deviations from the individual molecules in the previously reported monoclinic crystals are 0.50-0.57 A. This result proves that the structure of monellin is not significantly influenced by crystal packing forces.
- Macromolecular Structure Laboratory, NCI-Frederick Cancer Research and Development Center, ABL-Basic Research Program, MD 21702, USA.
Organizational Affiliation: