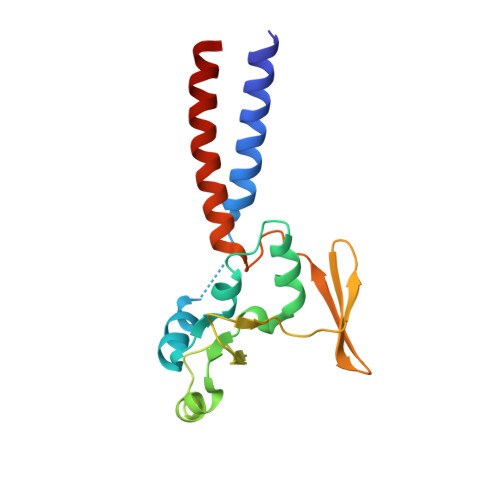
Structural basis for the MukB-topoisomerase IV interaction and its functional implications in vivo.
Vos, S.M., Stewart, N.K., Oakley, M.G., Berger, J.M.(2013) EMBO J 32: 2950-2962
- PubMed: 24097060 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2013.218
- Primary Citation Related Structures:
4MN4 - PubMed Abstract:
Chromosome partitioning in Escherichia coli is assisted by two interacting proteins, topoisomerase (topo) IV and MukB. MukB stimulates the relaxation of negative supercoils by topo IV; to understand the mechanism of their action and to define this functional interplay, we determined the crystal structure of a minimal MukB-topo IV complex to 2.3 Å resolution. The structure shows that the so-called 'hinge' region of MukB forms a heterotetrameric assembly with a C-terminal DNA binding domain (CTD) on topo IV's ParC subunit. Biochemical studies show that the hinge stimulates topo IV by competing for a site on the CTD that normally represses activity on negatively supercoiled DNA, while complementation tests using mutants implicated in the interaction reveal that the cellular dependency on topo IV derives from a joint need for both strand passage and MukB binding. Interestingly, the configuration of the MukB·topo IV complex sterically disfavours intradimeric interactions, indicating that the proteins may form oligomeric arrays with one another, and suggesting a framework by which MukB and topo IV may collaborate during daughter chromosome disentanglement.
- Department of Molecular and Cell Biology, University of California, Berkeley, CA, USA.
Organizational Affiliation: