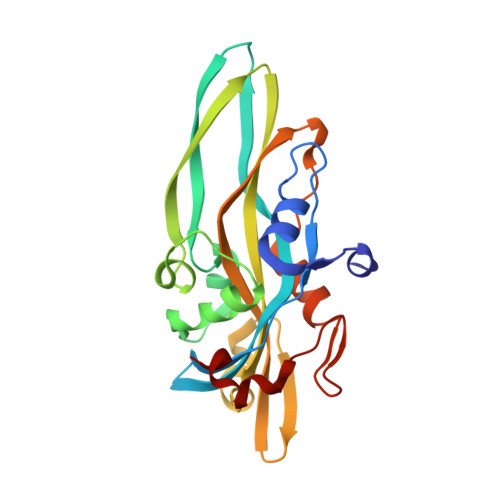
X-ray and Cryo-electron Microscopy Structures of Monalysin Pore-forming Toxin Reveal Multimerization of the Pro-form.
Leone, P., Bebeacua, C., Opota, O., Kellenberger, C., Klaholz, B., Orlov, I., Cambillau, C., Lemaitre, B., Roussel, A.(2015) J Biological Chem 290: 13191-13201
- PubMed: 25847242 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.646109
- Primary Citation Related Structures:
4MJT, 4MKO, 4MKQ - PubMed Abstract:
β-Barrel pore-forming toxins (β-PFT), a large family of bacterial toxins, are generally secreted as water-soluble monomers and can form oligomeric pores in membranes following proteolytic cleavage and interaction with cell surface receptors. Monalysin has been recently identified as a β-PFT that contributes to the virulence of Pseudomonas entomophila against Drosophila. It is secreted as a pro-protein that becomes active upon cleavage. Here we report the crystal and cryo-electron microscopy structure of the pro-form of Monalysin as well as the crystal structures of the cleaved form and of an inactive mutant lacking the membrane-spanning region. The overall structure of Monalysin displays an elongated shape, which resembles those of β-pore-forming toxins, such as Aerolysin, but is devoid of a receptor-binding domain. X-ray crystallography, cryo-electron microscopy, and light-scattering studies show that pro-Monalysin forms a stable doughnut-like 18-mer complex composed of two disk-shaped nonamers held together by N-terminal swapping of the pro-peptides. This observation is in contrast with the monomeric pro-form of the other β-PFTs that are receptor-dependent for membrane interaction. The membrane-spanning region of pro-Monalysin is fully buried in the center of the doughnut, suggesting that upon cleavage of pro-peptides, the two disk-shaped nonamers can, and have to, dissociate to leave the transmembrane segments free to deploy and lead to pore formation. In contrast with other toxins, the delivery of 18 subunits at once, nearby the cell surface, may be used to bypass the requirement of receptor-dependent concentration to reach the threshold for oligomerization into the pore-forming complex.
- From the CNRS, Architecture et Fonction des Macromolécules Biologiques (AFMB), Unité Mixte de Recherche (UMR) 7257, 13288 Marseille, France, the Aix-Marseille Université, Architecture et Fonction des Macromolécules Biologiques, UMR 7257, 13288 Marseille, France.
Organizational Affiliation: