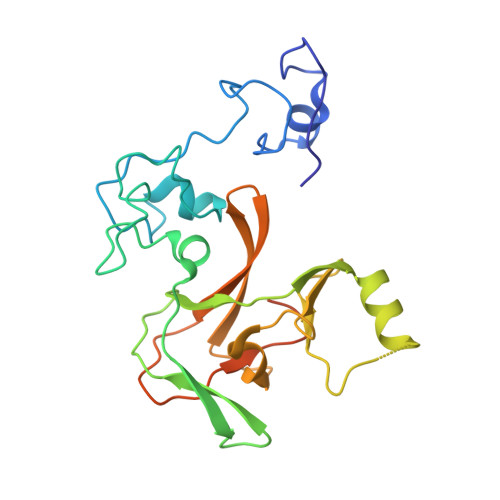
Structure of the catalytic domain of EZH2 reveals conformational plasticity in cofactor and substrate binding sites and explains oncogenic mutations.
Wu, H., Zeng, H., Dong, A., Li, F., He, H., Senisterra, G., Seitova, A., Duan, S., Brown, P.J., Vedadi, M., Arrowsmith, C.H., Schapira, M.(2013) PLoS One 8: e83737-e83737
- PubMed: 24367611 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0083737
- Primary Citation Related Structures:
4MI0 - PubMed Abstract:
Polycomb repressive complex 2 (PRC2) is an important regulator of cellular differentiation and cell type identity. Overexpression or activating mutations of EZH2, the catalytic component of the PRC2 complex, are linked to hyper-trimethylation of lysine 27 of histone H3 (H3K27me3) in many cancers. Potent EZH2 inhibitors that reduce levels of H3K27me3 kill mutant lymphoma cells and are efficacious in a mouse xenograft model of malignant rhabdoid tumors. Unlike most SET domain methyltransferases, EZH2 requires PRC2 components, SUZ12 and EED, for activity, but the mechanism by which catalysis is promoted in the PRC2 complex is unknown. We solved the 2.0 Å crystal structure of the EZH2 methyltransferase domain revealing that most of the canonical structural features of SET domain methyltransferase structures are conserved. The site of methyl transfer is in a catalytically competent state, and the structure clarifies the structural mechanism underlying oncogenic hyper-trimethylation of H3K27 in tumors harboring mutations at Y641 or A677. On the other hand, the I-SET and post-SET domains occupy atypical positions relative to the core SET domain resulting in incomplete formation of the cofactor binding site and occlusion of the substrate binding groove. A novel CXC domain N-terminal to the SET domain may contribute to the apparent inactive conformation. We propose that protein interactions within the PRC2 complex modulate the trajectory of the post-SET and I-SET domains of EZH2 in favor of a catalytically competent conformation.
- Structural Genomics Consortium, University of Toronto, Toronto, Ontario, Canada.
Organizational Affiliation: