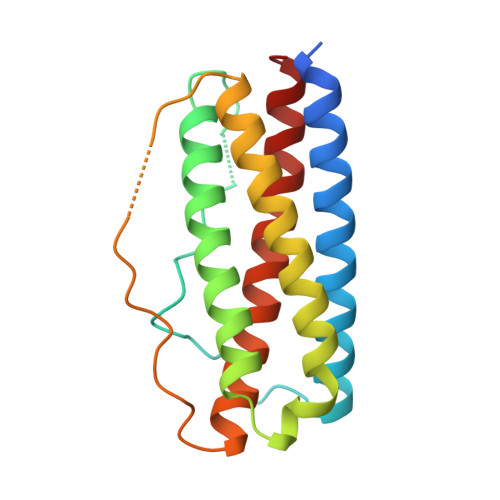
The structure of human interleukin-11 reveals receptor-binding site features and structural differences from interleukin-6.
Putoczki, T.L., Dobson, R.C., Griffin, M.D.(2014) Acta Crystallogr D Biol Crystallogr 70: 2277-2285
- PubMed: 25195742 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004714012267
- Primary Citation Related Structures:
4MHL - PubMed Abstract:
Interleukin (IL)-11 is a multifunctional member of the IL-6 family of cytokines. Recombinant human IL-11 is administered as a standard clinical treatment for chemotherapy-induced thrombocytopaenia. Recently, a new role for IL-11 signalling as a potent driver of gastrointestinal cancers has been identified, and it has been demonstrated to be a novel therapeutic target for these diseases. Here, the crystal structure of human IL-11 is reported and the structural resolution of residues previously identified as important for IL-11 activity is presented. While IL-11 is thought to signal via a complex analogous to that of IL-6, comparisons show important differences between the two cytokines and it is suggested that IL-11 engages GP130 differently to IL-6. In addition to providing a structural platform for further study of IL-11, these data offer insight into the binding interactions of IL-11 with each of its receptors and the structural mechanisms underlying agonist and antagonist variants of the protein.
- Inflammation Division, Walter and Eliza Hall Institute of Medical Research, Parkville, Victoria 3052, Australia.
Organizational Affiliation: