Structures of human DNA polymerase beta inserting bases opposite templating O6MG
Koag, M.C., Min, K., Monzingo, A.F., Lee, S.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA polymerase beta | 325 | Homo sapiens | Mutation(s): 0 Gene Names: POLB EC: 2.7.7.7 (PDB Primary Data), 4.2.99 (PDB Primary Data), 4.2.99.18 (UniProt) | 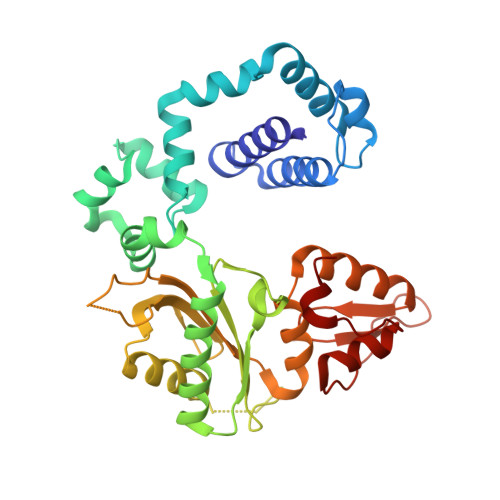 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P06746 GTEx: ENSG00000070501 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P06746 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| template | B [auth T] | 16 | N/A | 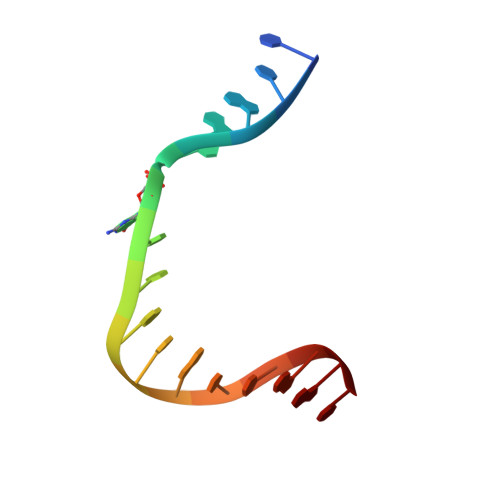 |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 3 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| up primer | C [auth P] | 11 | N/A | 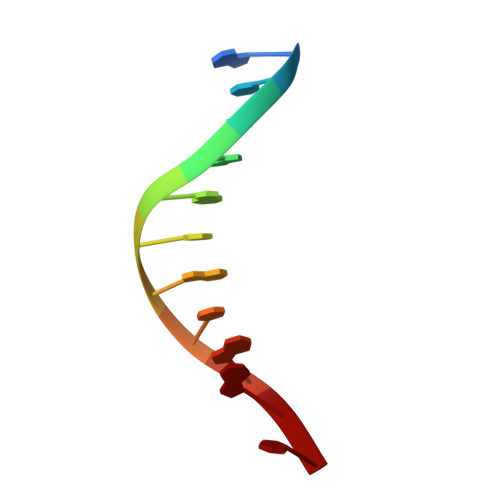 |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PO4 Download:Ideal Coordinates CCD File | H [auth A] | PHOSPHATE ION O4 P NBIIXXVUZAFLBC-UHFFFAOYSA-K |  | ||
| MG Download:Ideal Coordinates CCD File | F [auth A], I [auth P] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| NA Download:Ideal Coordinates CCD File | E [auth A], G [auth A] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 54.971 | α = 90 |
| b = 79.662 | β = 106.68 |
| c = 55.001 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| StructureStudio | data collection |
| DENZO | data reduction |
| HKL-2000 | data scaling |
| MOLREP | phasing |