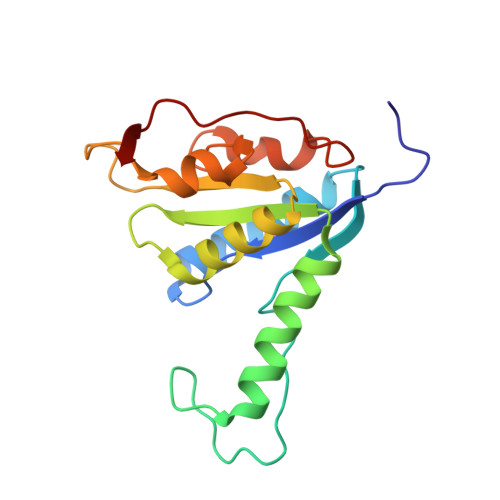
Glycosylation of a tricyclic purine analog at alternative sites by nucleoside 2 -deoxyribosyltransferases
Ye, W., Paul, D., Gao, L., Seckute, J., Sangaiah, R., Karupiah, J., Zhang, Z., Kaminski, A.P., Ealick, S.E., Gold, A., Ball, L.M.(2014) PLoS One