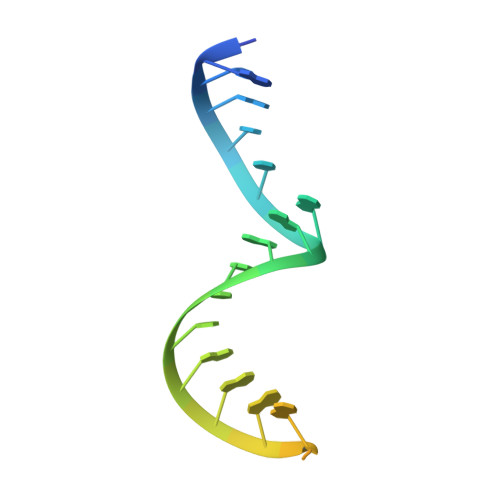
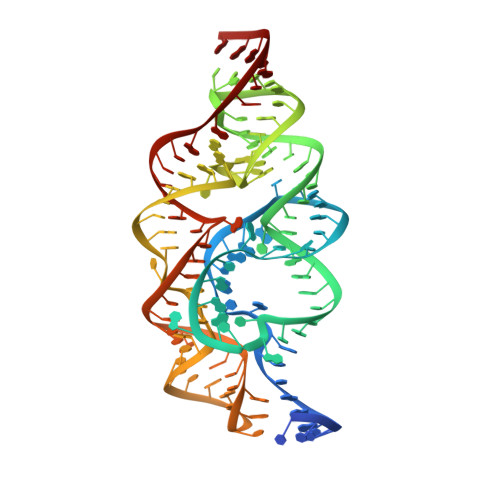
An in vitro evolved glmS ribozyme has the wild-type fold but loses coenzyme dependence.
Lau, M.W., Ferre-D'Amare, A.R.(2013) Nat Chem Biol 9: 805-810
- PubMed: 24096303 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.1360
- Primary Citation Related Structures:
4MEG, 4MEH - PubMed Abstract:
Uniquely among known ribozymes, the glmS ribozyme-riboswitch requires a small-molecule coenzyme, glucosamine-6-phosphate (GlcN6P). Although consistent with its gene-regulatory function, the use of GlcN6P is unexpected because all of the other characterized self-cleaving ribozymes use RNA functional groups or divalent cations for catalysis. To determine what active site features make this ribozyme reliant on GlcN6P and to evaluate whether it might have evolved from a coenzyme-independent ancestor, we isolated a GlcN6P-independent variant through in vitro selection. Three active site mutations suffice to generate a highly reactive RNA that adopts the wild-type fold but uses divalent cations for catalysis and is insensitive to GlcN6P. Biochemical and crystallographic comparisons of wild-type and mutant ribozymes show that a handful of functional groups fine-tune the RNA to be either coenzyme or cation dependent. These results indicate that a few mutations can confer new biochemical activities on structured RNAs. Thus, families of structurally related ribozymes with divergent function may exist.
- National Heart, Lung and Blood Institute, Bethesda, Maryland, USA.
Organizational Affiliation: