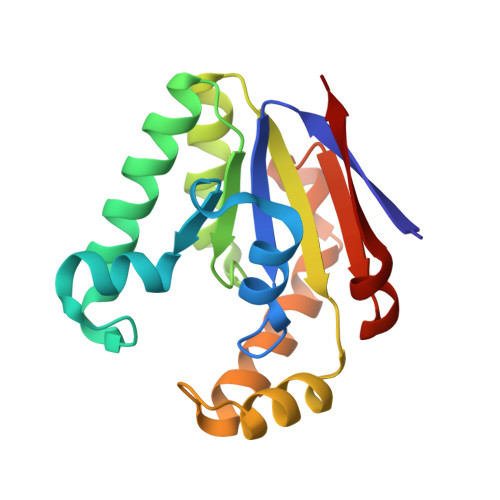
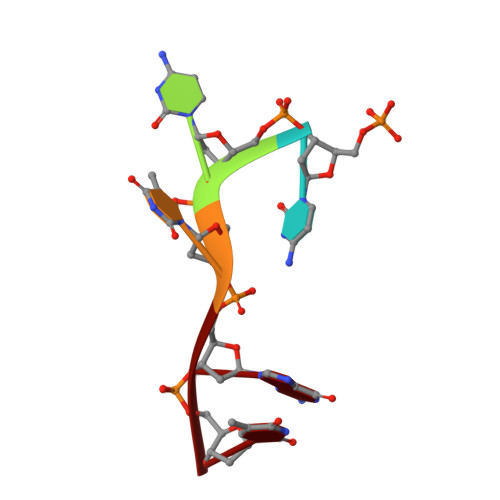
Structures of bacterial polynucleotide kinase in a Michaelis complex with GTP*Mg2+ and 5'-OH oligonucleotide and a product complex with GDP*Mg2+ and 5'-PO4 oligonucleotide reveal a mechanism of general acid-base catalysis and the determinants of phosphoacceptor recognition.
Das, U., Wang, L.K., Smith, P., Jacewicz, A., Shuman, S.(2014) Nucleic Acids Res 42: 1152-1161
- PubMed: 24150947 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkt936
- Primary Citation Related Structures:
4MDE, 4MDF - PubMed Abstract:
Clostridium thermocellum polynucleotide kinase (CthPnk), the 5' end-healing module of a bacterial RNA repair system, catalyzes reversible phosphoryl transfer from an NTP donor to a 5'-OH polynucleotide acceptor. Here we report the crystal structures of CthPnk-D38N in a Michaelis complex with GTP•Mg(2+) and a 5'-OH oligonucleotide and a product complex with GDP•Mg(2+) and a 5'-PO4 oligonucleotide. The O5' nucleophile is situated 3.0 Å from the GTP γ phosphorus in the Michaelis complex, where it is coordinated by Asn38 and is apical to the bridging β phosphate oxygen of the GDP leaving group. In the product complex, the transferred phosphate has undergone stereochemical inversion and Asn38 coordinates the 5'-bridging phosphate oxygen of the oligonucleotide. The D38N enzyme is poised for catalysis, but cannot execute because it lacks Asp38-hereby implicated as the essential general base catalyst that abstracts a proton from the 5'-OH during the kinase reaction. Asp38 serves as a general acid catalyst during the 'reverse kinase' reaction by donating a proton to the O5' leaving group of the 5'-PO4 strand. The acceptor strand binding mode of CthPnk is distinct from that of bacteriophage T4 Pnk.
- Molecular Biology Program, Sloan-Kettering Institute, New York, NY 10065, USA.
Organizational Affiliation: