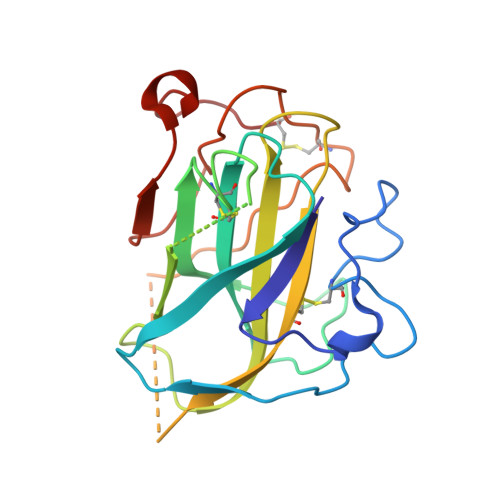
Discovery and characterization of a new family of lytic polysaccharide monooxygenases.
Hemsworth, G.R., Henrissat, B., Davies, G.J., Walton, P.H.(2014) Nat Chem Biol 10: 122-126
- PubMed: 24362702 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.1417
- Primary Citation Related Structures:
4MAH, 4MAI - PubMed Abstract:
Lytic polysaccharide monooxygenases (LPMOs) are a recently discovered class of enzymes capable of oxidizing recalcitrant polysaccharides. They are attracting considerable attention owing to their potential use in biomass conversion, notably in the production of biofuels. Previous studies have identified two discrete sequence-based families of these enzymes termed AA9 (formerly GH61) and AA10 (formerly CBM33). Here, we report the discovery of a third family of LPMOs. Using a chitin-degrading exemplar from Aspergillus oryzae, we show that the three-dimensional structure of the enzyme shares some features of the previous two classes of LPMOs, including a copper active center featuring the 'histidine brace' active site, but is distinct in terms of its active site details and its EPR spectroscopy. The newly characterized AA11 family expands the LPMO clan, potentially broadening both the range of potential substrates and the types of reactive copper-oxygen species formed at the active site of LPMOs.
- Department of Chemistry, University of York, Heslington, York, UK.
Organizational Affiliation: